Advertisement
Review Series
Open Access |  10.1172/JCI199845
10.1172/JCI199845
Lysosomal homeostasis at the crossroads of neurodegeneration
Stefano De Tito1 and Sharon A. Tooze2
1Navira Bio at
2The Francis Crick Institute, London, United Kingdom.
Address correspondence to Stefano De Tito, Navira Bio, The Francis Crick Institute, 1 Midland Road, London, W1 AT, United Kingdom. Email: stefano.detito@navira.bio. Or to: Sharon A. Tooze, The Francis Crick Institute, 1 Midland Road, London W1 1AT, United Kingdom. Email: Sharon.tooze@crick.ac.uk.
Find articles by De Tito, S. in: PubMed | Google Scholar
1Navira Bio at
2The Francis Crick Institute, London, United Kingdom.
Address correspondence to Stefano De Tito, Navira Bio, The Francis Crick Institute, 1 Midland Road, London, W1 AT, United Kingdom. Email: stefano.detito@navira.bio. Or to: Sharon A. Tooze, The Francis Crick Institute, 1 Midland Road, London W1 1AT, United Kingdom. Email: Sharon.tooze@crick.ac.uk.
Find articles by
Tooze, S.
in:
PubMed
|
Google Scholar
|

Published April 1, 2026 - More info
J Clin Invest. 2026;136(7):e199845. https://doi.org/10.1172/JCI199845.
© 2026 De Tito et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
-
Abstract
Lysosomes function as metabolic control centers that integrate degradation, nutrient sensing, and stress signaling. In neurons, which must maintain proteostasis and energetic balance throughout life, lysosomal homeostasis determines cellular resilience. Emerging evidence identifies lysosomal injury and defective repair as common denominators across neurodegenerative diseases. Damage to the lysosomal membrane caused by oxidative stress, lipid imbalance, or genetic mutations triggers a hierarchical quality control cascade. Early lesions recruit the endosomal sorting complex required for transport (ESCRT) machinery for mechanical resealing, while larger ruptures activate lipid-centered recovery modules. When repair fails, lysophagy eliminates irreparable organelles and a TFEB-dependent transcriptional program regenerates the lysosomal pool. These tightly coupled responses safeguard neurons from catastrophic proteostatic collapse. Their impairment, through mutations in lysosomal proteins, or through aging, produces the lysosomal fragility that underlies Alzheimer disease, Parkinson disease, amyotrophic lateral sclerosis/frontotemporal dementia, and Huntington disease. Crosstalk between lysosomes, mitochondria, and ER integrates local damage with systemic metabolic adaptation, while dysregulated lysosomal exocytosis and inflammation propagate pathology. Understanding how ESCRT complexes, lipid transport, and transcriptional renewal cooperate to preserve lysosomal integrity reveals unifying principles of neurodegeneration and defines molecular targets for intervention. Restoring lysosomal repair and renewal offers a rational path toward preventing neuronal loss.
-
Introduction
The lysosome, an acidic membrane-bound organelle rich in hydrolytic enzymes, is now recognized as a dynamic signaling hub central to metabolism, quality control, stress adaptation, and inter-organelle communication (1–6). The membrane of the lysosome anchors key signaling complexes, including mammalian target of rapamycin complex 1 (mTORC1), AMP-activated protein kinase (AMPK), and transcription factor EB (TFEB), which link nutrient and energy status to global cellular programs (7, 8).
In neurons, lysosomal homeostasis is uniquely critical due to their postmitotic nature and extreme polarization (9, 10). Maintaining lysosomal acidity, enzyme content, and trafficking fidelity across long axons is an ongoing challenge. Neuronal lysosomes not only degrade cargo but also mediate synaptic remodeling and plasma membrane repair (11–13). Even subtle defects in lysosomal function can initiate progressive proteostatic and metabolic failure that culminates in neurodegeneration.
Genetic and pathological evidence firmly places lysosomal dysfunction at the center of neuronal loss (14–17). Recent lysosome immunopurification (Lyso-IP) studies revealed disease-associated accumulation of aggregation-prone proteins such as α-synuclein, tau, and TDP-43 that precedes overt membrane damage (18, 19). Mutations in lysosomal proteins that disrupt hydrolase trafficking, affect lysosomal membrane repair, or alter lipid metabolism are implicated in diverse neurodegenerative diseases (Table 1). In sporadic conditions, neurodegeneration, oxidative stress, lipid peroxidation, and chronic mTORC1 activation progressively weaken lysosomal integrity (10, 20–22).
When the lysosomal membrane becomes damaged and leaky, a process termed lysosomal membrane permeabilization (LMP), cells activate a hierarchical lysosomal quality control system (23, 24). Depending on the extent of damage, this system coordinates mechanical repair, lipid remodeling, selective clearance, and transcriptional regeneration (24–26). Successful recovery restores homeostasis, whereas failure results in inflammasome activation, mitochondrial dysfunction, and neuronal death (6, 10, 27, 28).
-
Mechanisms of lysosomal damage and repair
LMP occurs along a continuum from transient nanoscopic pores to catastrophic rupture (29–31). Even limited permeabilization exposes luminal glycans to the cytosol, rapidly recruiting galectins-3, -8, and -9 (32–35). Galectins are a conserved family of cytosolic lectins containing carbohydrate recognition domains that bind β-galactoside–containing glycoconjugates. As most lysosomal membrane proteins are glycosylated, membrane disruption exposes their β-galactosides to the cytosol, where they are rapidly detected by galectins that serve as sentinels of lysosomal integrity (34). Galectin-3 interacts with the endosomal sorting complex required for transport (ESCRT) adaptor ALIX and TSG101 to nucleate ESCRT-III filaments that constrict and seal the membrane leak (33, 36). Local calcium efflux through the lesion activates ALG-2, which bridges ALIX to phospholipids and stabilizes ESCRT polymerization (37–39) (Figure 1). The AAA ATPase VPS4 subsequently disassembles ESCRT-III, recycling the subunits for additional rounds of repair (40, 41). In neurons, this process unfolds within seconds, preserving lysosomal integrity and preventing the leakage of cathepsins that would otherwise activate inflammasomes and induce mitochondrial depolarization (42, 43).
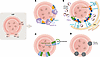 Figure 1
Figure 1Mechanisms of lysosomal membrane repair. (A) The ESCRT machinery, recruited by galectin-3 (Gal3) and ALIX, polymerizes at rupture sites to reseal small pores. (B) The PI4K2A/ORP/ATG9A lipid-transfer axis forms ER-lysosome bridges that restore membrane lipid composition and curvature. ATG2 cooperates with ATG9A vesicles to deliver lipids for bilayer expansion. (C) Neutral sphingomyelinase–dependent conversion of sphingomyelin to ceramide promotes local fusion and sealing. (D) Under severe stress, protein-RNA condensates assemble into transient stress granule patches that shield lesions until structural repair completes. Together, these coordinated modules preserve lysosomal integrity and prevent cathepsin release.
Concurrently, galectin sensors transduce metabolic signals that transiently suppress growth programs and promote autophagy. Galectin-8 assembles the GALTOR complex, which binds the Ragulator/Rag axis and inhibits mTORC1, thereby facilitating TFEB activation (34, 35). Galectin-9, through USP9X-dependent TAK1 ubiquitination, stimulates AMPK and broad autophagic responses (44, 45). Together, these events couple structural repair with metabolic reprogramming, ensuring that the cell prioritizes restoration over growth during lysosomal stress.
When membrane injury exceeds ESCRT capacity, a lipid-centered recovery module is engaged (46–50). Phosphatidylinositol 4-kinase type 2-α (PI4K2A) generates phosphatidylinositol-4-phosphate (PI4P) at the damaged surface, stimulating the phosphoinositide-initiated tethering and lipid-transport (PITT) pathway (48, 49) (Figure 1). PI4P recruits oxysterol-binding protein (OSBP) and the OSBP-related proteins (ORPs) ORP9, ORP10, and ORP11, which tether the lysosome to VAPA and VAPB on the ER, establishing a lipid-exchange bridge. Through these contacts, PI4P is transferred to the ER, while phosphatidylserine (PS) and cholesterol flow back to the lysosome. Furthermore, the lipid transfer protein ATG2 facilitates phospholipid flow from the ER to damaged lysosomes, promoting membrane expansion and restoration (49, 51). This bidirectional transfer restores bilayer asymmetry, curvature, and mechanical resilience. A pivotal element of this process is ATG9A, a transmembrane protein essential for autophagy (46, 52–55). ATG9A mainly resides in the Golgi and endosomes and cycles between other organelles, including the plasma membrane and lysosome (46, 50, 56). It has been shown to have lipid scramblase activity that is required for its function in autophagy (53, 55) and membrane repair (46, 57). ATG9A traffics from the trans-Golgi network to the damaged lysosome in response to injury (46, 50). ATG9A vesicles deliver PI4K2A to the lesion site, where newly synthesized PI4P creates microdomains to seal lysosomal membrane damage via lipid transfer of ORPs through ER-lysosome contact sites (46) (Figure 1). ARFIP2, a Bin/amphiphysin/Rvs homology (BAR) domain–containing protein that detects membrane curvature (58, 59), senses these PI4P-enriched zones (60), coordinating the lipid-transfer proteins to complete the resealing process. This ATG9A/PI4K2A/ARFIP2 axis has been defined as an autonomous repair route parallel to ESCRT. However, whether ESCRT- and lipid-mediated repair pathways operate sequentially, competitively, or in a damage-specific hierarchy remains an open question. Disruption of ATG9A trafficking or ARFIP2 sensing leads to alteration in lysosomal homeostasis, a phenotype similar to that observed altering vesicle trafficking in dopaminergic neurons of Parkinson disease (PD) models (61–63).
Sphingolipid metabolism further contributes to membrane recovery. Activation of sphingomyelin scramblases exposes sphingomyelin to the cytosolic leaflet, where neutral sphingomyelinase generates ceramide (64). Ceramide’s conical geometry promotes local fusion and sealing, while cholesterol imported from the ER through OSBP stiffens the membrane and reduces permeability (64–66). The interplay among PI4P, PS, ceramide, and cholesterol defines a lipid code that determines whether a damaged lysosome is repaired or removed (26) (Figure 1).
In parallel, recent evidence suggests that stress granules transiently associate with damaged lysosomes and may supply membrane components that limit leakage (67) (Figure 1). This stress granule–mediated sealing provides an auxiliary repair route that complements ESCRT- and lipid-driven mechanisms, particularly during acute proteotoxic stress.
Finally, these mechanisms of lysosomal repair are also reinforced by cytoskeletal remodeling. Following LMP, leucine-rich-repeat kinase 2 (LRRK2) is recruited by RAB29 and phosphorylates RAB8A and RAB10, initiating JIP4-dependent tubulation (61–63, 68). Actin nucleation through the actin-related protein 2/3 complex subunit 2 (ARPC2) and formin-1 maintains curvature and mechanical tension until lipid reconstitution is complete (68). Pathogenic LRRK2 mutations disrupt this choreography and lead to defective repair and dopaminergic neuron vulnerability (61–63).
-
Autophagy and lysophagy
To sustain cellular homeostasis and promote survival under stress conditions, eukaryotic cells depend on autophagy, a conserved catabolic pathway that delivers cytoplasmic material to lysosomes for degradation. During this process, portions of cytoplasm, damaged organelles, misfolded proteins, and invading pathogens are enclosed within double-membraned structures termed autophagosomes (69). These form de novo from the ER (70). A defining event in autophagosome formation is the emergence of the phagophore, a crescent-shaped isolation membrane whose molecular assembly remains only partially understood. As the phagophore expands (71), it engulfs cytoplasmic material marked for turnover and eventually seals to generate a mature autophagosome (69). The completed autophagosome then fuses sequentially with late endosomes and lysosomes, where its contents are degraded and recycled to sustain cellular metabolism. This process is choreographed by a highly conserved set of autophagy-related (ATG) proteins that coordinate each stage of autophagosome initiation, maturation, and fusion with the lysosomal compartment (69, 72) (Table 2 and Figure 2).
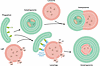 Figure 2
Figure 2Autophagy and lysophagy pathways. Schematic overview of canonical autophagy and selective lysophagy. Under basal or stress conditions, cytoplasmic material, damaged organelles, and protein aggregates are sequestered by expanding phagophores to form double-membrane autophagosomes. These vesicles fuse sequentially with late endosomes and lysosomes for degradation and recycling. In lysophagy, LMP exposes luminal glycans that recruit galectins, ubiquitin ligases, and autophagy receptors. The phagophore assembly machinery then encapsulates damaged lysosomes for removal. Successful completion restores lysosomal homeostasis, while failure triggers inflammasome activation and neurodegeneration.
 Table 2
Table 2Autophagy-related (ATG) proteins and complexes involved in autophagosome biogenesis and maturation
In selective autophagy, specific cellular components are identified and modified by E3 ubiquitin ligases, which conjugate ubiquitin, a small regulatory protein that signals for degradation, to selected substrates (73, 74). These ubiquitinated cargos are recognized by autophagy adaptor proteins that link them to the autophagic machinery (75). In lysophagy, the selective removal of damaged lysosomes, membrane rupture triggers the accumulation of galectins and ubiquitin on the lysosomal surface (25, 76, 77) (Figure 2). Upon LMP, galectins bind the exposed β-galactosides (33, 34). In parallel, ubiquitination provides a complementary signal for the recognition of damaged lysosomes (73, 78). Following rupture, lysosome-associated membrane protein 1 (LAMP1), and 2 (LAMP2), and transmembrane protein 192 (TMEM192) are modified by ubiquitin (75, 77, 79). These modifications are catalyzed by distinct enzyme complexes, including the SCFFBXO27 E3 ligase complex, composed of S-phase–kinase–associated protein 1 (SKP1), cullin-1 (CUL1), and F-box protein 27 (FBXO27) (80), and the cullin-4A–DNA damage–binding protein 1–WD repeat and FYVE domain–containing 1 (CUL4A-DDB1-WDFY1) complex (81). The ubiquitin-conjugating enzyme E2 Q-like 1 (UBE2QL1) also collaborates with lysophagy effectors to promote LAMP1 ubiquitination (78). Once the ubiquitinated and galectin-decorated lysosomal membrane is identified, the phagophore assembly machinery is rapidly mobilized to initiate encapsulation. Galectin-3 and TRIM16 form a signaling platform that recruits the ULK1-ATG13-FIP200 initiation complex (Table 2), marking the site for autophagosome nucleation (82). Consistent with this organizer role, galectin-enriched microdomains help concentrate ubiquitin signals and LC3-binding adaptors, enabling robust lysophagy even when damage is spatially restricted. Specifically, galectin-8 engages the autophagy receptor NDP52, which bridges damaged lysosomes to the microtubule-associated protein 1 light chain 3–positive (LC3-positive) membranes (83). Ubiquitin-binding adaptors, including sequestosome 1 (SQSTM1, also known as p62), tax1-binding protein 1 (TAX1BP1), optineurin (OPTN), and calcium-binding and coiled-coil domain–containing 2 (CALCOCO2, also known as NDP52L) cluster at the lesion, tethering the expanding phagophore to the lysosomal surface (77, 84–86). These adaptors, through their LC3-interacting regions, recruit the ATG5-ATG12-ATG16L1 conjugation system to drive LC3 lipidation and elongation of the isolation membrane around the damaged organelle (87). As the phagophore closes, chaperones such as heat shock protein family B (small) member 1 (HSPB1) and the segregase valosin-containing protein (VCP, also known as p97) remove ubiquitinated membrane remnants, ensuring efficient engulfment (88). The mature autophagosome then fuses with functional lysosomes for degradation of the damaged compartment. This coordinated sequence restores lysosomal homeostasis (25). Recent work also identifies SPART (also known as SPG20) as a lysosomal quality control factor that promotes lysophagy of damaged lysosomes by coordinating early damage recognition with downstream recruitment of autophagy machinery (76). Yet, how cells distinguish repairable from irreversibly damaged lysosomes and how this decision changes with age remains incompletely understood.
After elimination of irreparable organelles, cells restore lysosomal mass through transcriptional regeneration. Calcium efflux through transient receptor potential mucolipin-1 (TRPML1, also known as MCOLN1) on the lysosome membrane activates calcineurin, which dephosphorylates TFEB to drive nuclear translocation and expression of lysosomal and autophagy genes (89). SMAD-specific E3 ubiquitin protein ligase 1–mediated (SMURF1-mediated) calcineurin activation further amplifies TFEB signaling (90). Nuclear TFEB upregulates synthesis of enzymes, transporters, lipid-metabolizing proteins, and factors controlling lysosomal exocytosis, a Ca²+-dependent process in which lysosomes fuse with the plasma membrane to release their contents and supply membrane for repair and clearance (7, 89). Whether prolonged TFEB activation remains protective or becomes maladaptive under chronic stress conditions is still debated. In neurons, this renewal pathway replenishes distal lysosomes critical for synaptic maintenance (91, 92). Failure of this regenerative response results in persistent leakage, chronic activation of the NOD LRR- and pyrin domain–containing 3 (NLRP3) and cyclic GMP-AMP synthase (cGAS)–stimulator of interferon gene (STING) pathways, and progressive neuroinflammation (93). Mechanistically, lysosomal damage can promote cytosolic DNA signaling (via mitochondrial or nuclear DNA leakage) to activate cGAS/STING, while STING itself has been proposed to localize to endolysosomal membranes and directly influence lysosomal recovery programs, including TFEB-dependent transcription.
-
Lysosomal dysfunction in neurodegenerative diseases
Lysosomal dysfunction represents a common axis of vulnerability across major neurodegenerative disorders (Table 1), where it both precedes and amplifies the accumulation of toxic proteins (17, 23, 51). While extensive evidence supports a role for lysosomal dysfunction in neurodegeneration, the specific contribution of membrane repair and lysosomal quality control pathways to disease initiation versus progression remains an active area of investigation. Most studies to date have documented broad defects in lysosomal degradation, trafficking, or acidification, whereas direct evidence indicating that failure of specific membrane repair pathways such as ESCRT-, PITT-, or lysophagy-mediated mechanisms is causal of disease is still emerging.
When lysosomal damage becomes chronic or exceeds the capacity for repair and lysophagy, it can activate regulated cell death pathways that directly contribute to neuronal loss (10). Extensive lysosomal membrane permeabilization allows the cytosolic release of cathepsins, redox-active iron, and other hydrolases that trigger mitochondrial dysfunction, caspase activation, lipid peroxidation, and inflammatory signaling. Depending on cellular context, these events can engage apoptosis, necroptosis, pyroptosis, or ferroptosis, positioning lysosomes as central hubs that link metabolic stress to cell fate decisions (20–22). In neurons, which rely heavily on lysosomal integrity for long-term proteostasis, persistent lysosomal leakage lowers the threshold for maladaptive cell death, thereby providing a mechanistic bridge between organelle dysfunction and progressive neurodegeneration.
The inability to maintain lysosomal integrity, acidification, or repair capacity directly correlates with neuronal loss in Alzheimer disease (AD) (43), PD (94), amyotrophic lateral sclerosis/frontotemporal dementia (ALS/FTD) (20), and Huntington disease (HD) (95). These findings suggest that impaired lysosomal repair, replacement, and regeneration pathways may act as important modulators of neuronal resilience rather than sole drivers of neurodegenerative pathology.
AD. In AD, lysosomal impairment arises early, often preceding the extracellular accumulation of amyloid-β (Aβ) peptides and the intracellular aggregation of tau, a microtubule-associated protein that forms neurofibrillary tangles. Aβ peptides are generated by sequential cleavage of amyloid precursor protein (APP), a transmembrane protein involved in neuronal growth and synaptic repair, by β- and γ-secretases (96, 97). Mutations in presenilin 1 (PSEN1) and 2 (PSEN2), core components of the γ-secretase complex, alter APP processing and disrupt assembly of the lysosomal vacuolar ATPase (V-ATPase), impairing endolysosomal acidification, hydrolase maturation, and proteolytic efficiency (98–105).
Aβ and tau pathology also compromise the blood-brain barrier (BBB), increasing its permeability to apolipoprotein B–bound (ApoB-bound) cholesterol, which is typically restricted to the peripheral circulation (106, 107). The influx of peripheral cholesterol into the brain perturbs neuronal endolysosomal structure and function, producing enlarged, deacidified lysosomes with reduced degradative capacity (108, 109).
Furthermore, apolipoprotein E (ApoE), particularly the ε4 isoform, modulates lipid trafficking and lysosomal proteolysis in glial cells, further contributing to impaired clearance and neuroinflammation (110–112).
Another hallmark of AD is the accumulation of cathepsin-positive but incompletely digested autophagic vacuoles within dystrophic neurites, indicating inefficient fusion and maturation of autophagosomes into degradative lysosomes (113, 114).
Tau fibrils internalized by neurons and astrocytes accumulate within lysosomes, causing swelling, deacidification, and recruitment of ESCRT machinery; this nanoscale membrane injury promotes cytosolic tau nucleation at the lysosomal surface, suggesting that lysosomal damage may actively initiate tau aggregation (115).
Persistent mTORC1 activity and impaired TFEB/transcription factor binding to IGHM enhancer 3 (TFE3) signaling in AD further suppress lysosomal biogenesis and clearance (116, 117), while defective axonal trafficking causes lysosome accumulation in dystrophic neurites, amplifying local proteostatic collapse (11).
In addition, APP C-terminal fragments have been reported to directly perturb endolysosomal/lysosomal physiology, suggesting that APP-derived species can drive lysosomal stress upstream of overt plaque pathology (118).
Finally, elevated ceramide species in human AD cortices indicate persistent activation of lipid-based repair pathways, underscoring lysosomal injury as an initiating lesion rather than a downstream consequence (119).
At the mechanistic level, these changes are consistent with reduced ESCRT-mediated membrane repair capacity, impaired TFEB-dependent lysosomal renewal, and chronic lipid-driven membrane stress that may overwhelm may overwhelm membrane repair pathways involving the conversion of sphingomyelin to ceramide (sphingomyelin–ceramide).
PD. Lysosomal failure is both genetically and mechanistically central to the pathogenesis of PD (94, 120). Mutations in GBA1, which encodes the lysosomal enzyme β-glucocerebrosidase (GCase) responsible for hydrolyzing glucosylceramide into glucose and ceramide, reduce enzymatic activity and lead to the accumulation of glucosylceramide and related glycolipids (121–123). These lipid deposits destabilize lysosomal membranes and impair protease function. The resulting lipid imbalance promotes the aggregation of α-synuclein (encoded by SNCA), a presynaptic protein that normally regulates synaptic vesicle trafficking but, when misfolded, accumulates in lysosomes and further disrupts autophagic flux (124–126). Excess α-synuclein interferes with SNARE-mediated autophagosome-lysosome fusion, exacerbating autophagic inefficiency (124), while lysosomal exocytosis of α-synuclein aggregates contributes to extracellular propagation of pathology (127).
Further genetic evidence includes the finding that pathogenic mutations in LRRK2, which encodes a large multidomain kinase that phosphorylates Rab GTPases involved in vesicle trafficking, alter endolysosomal transport and lysosomal repair dynamics (61–63). Loss of ATP13A2 (PARK9), a lysosomal P-type ATPase that regulates polyamine and metal ion homeostasis, increases oxidative stress and impairs autophagic clearance (128–130). Finally, defective lysosomal acidification caused by mutations in transmembrane protein 175 (TMEM175), a lysosomal potassium/proton channel that maintains luminal pH, and vacuolar protein sorting-associated protein 35 (VPS35), a core component of the retromer complex that mediates cargo recycling, further compromises hydrolase activity and enzyme trafficking (131–137).
Together, these events create a feed-forward cycle of defective degradation and neuroinflammation driving neuronal vulnerability. These defects are consistent with impaired lipid-dependent membrane repair, including sphingomyelin–ceramide turnover and ATG9A-associated lipid exchange pathways, as well as altered lysophagy efficiency and Rab-regulated repair dynamics.
ALS and FTD. In ALS and FTD, diverse genetic mutations converge on lysosomal and autophagic dysfunction as a central pathogenic mechanism (17, 138). Progranulin (encoded by GRN), a secreted glycoprotein that is processed into granulin peptides within lysosomes, regulates lysosomal acidification, hydrolase trafficking, and membrane stability. Loss-of-function mutations in GRN lead to progranulin deficiency, which reduces lysosomal acidity and cathepsin activity, promoting the accumulation of ubiquitinated protein aggregates in the cytoplasm of neurons and glial cells (139–141). Hexanucleotide repeat expansions in chromosome 9 open reading frame 72 (C9orf72), the most common genetic cause of both ALS and FTD, cause two major pathogenic outcomes. First, decreased levels of the C9orf72 protein, which normally forms a complex with SMCR8 and WD repeat domain 41 (WDR41) to control endosomal trafficking and autophagy initiation, impair s lysosomal biogenesis and vesicle maturation (142). Second, repeat-associated non-AUG (RAN) translation of the expanded GGGGCC repeats generates toxic dipeptide repeat proteins (DPRs), including poly(GA), poly(GR), and poly(PR) that accumulate in the cytoplasm and nucleus. These DPRs disrupt lysosomal membranes, inhibit proteasome function, and interfere with nucleocytoplasmic transport by binding nuclear pore components (143–146).
Additional ALS/FTD-linked mutations further highlight the centrality of the autophagy-lysosome pathway. Mutations in VCP, an AAA+ ATPase that extracts ubiquitinated substrates from membranes and facilitates autophagosome maturation, impair degradation of damaged organelles (147). Changes in charged multivesicular body protein 2B (CHMP2B), a component of ESCR7-III, interfere with endosomal sorting and lysosomal membrane repair (148–150). Loss-of-function variants in TANK-binding kinase 1 (TBK1), a serine/threonine kinase that phosphorylates autophagy receptors OPTN and SQSTM1, weaken selective autophagy and the clearance of damaged mitochondria and lysosomes (151–153). Persistent lysosomal stress in ALS/FTD models induces chronic activation of the transcription factors TFEB and TFE3, which drive lysosomal and autophagy gene expression as a compensatory response to degradation failure, although whether prolonged activation becomes maladaptive remains under investigation (154). Furthermore, increased galectin-3 levels in patients with FTD highlight its prominent role in neuroinflammatory signaling that may influence disease pathogenesis (155).
Together, these events create a feed-forward cycle of defective degradation and neuroinflammation driving neuronal vulnerability. Several ALS/FTD-linked mutations, particularly those affecting ESCRT components and autophagy regulators, suggest a convergence on impaired membrane repair, inefficient lysophagy, and dysregulated TFEB/TFE3-mediated transcriptional adaptation.
HD. HD, caused by a CAG trinucleotide repeat expansion in the huntingtin gene (HTT), is characterized by progressive motor, cognitive, and psychiatric decline (156, 157). The mutant huntingtin protein (mHTT) contains an expanded polyglutamine (polyQ) tract that confers toxic gain-of-function properties and disrupts multiple facets of the lysosome-autophagosome network (95, 158). Under physiological conditions, wild-type HTT scaffolds autophagy and vesicular transport by interacting with HTT-associated protein 1 (HAP1), RAB7, and dynein, thereby mediating retrograde trafficking of autophagosomes and lysosomes along microtubules (159, 160). The mutant form impairs these interactions, leading to defective autophagosome recognition and inefficient transport of lysosomes to distal neuronal compartments, where degradation demand is highest (161, 162). mHTT also perturbs mTORC1 signaling at the lysosomal membrane, locking mTORC1 in an inactive configuration that suppresses TFEB activation and lysosomal gene transcription (163, 164).
Perturbations in lipid composition further compromise lysosomal membrane integrity and repair. In HD, phospholipid and sphingolipid composition is altered (including defective de novo sphingolipid synthesis) and cholesterol homeostasis is dysregulated, with accompanying changes in membrane fluidity; such lipid shifts are expected to stiffen membranes (e.g., via cholesterol/ceramide enrichment) and impede curvature-dependent lysosomal fusion/repair (165–169).
Moreover, mHTT interferes with the activity of RAB5 and RAB7, small GTPases that regulate endosome-lysosome maturation and fusion, resulting in stalled autophagic flux (159, 170).
Recent HD studies further link lysosomal dysfunction to altered galectin-3 signaling and dysregulation of TFEB/TFE3-controlled lysosomal programs, supporting lysosome-centered stress responses as disease modifiers (95, 171, 172).
These changes are consistent with compromised lysosomal renewal and defective recovery from membrane stress, potentially involving reduced TFEB activation, impaired lysophagy, and lipid composition changes that weaken sphingolipid-dependent repair capacity.
Lysosomal storage disorders. Lysosomal storage disorders (LSDs) comprise a group of more than 70 inherited disorders caused by mutations affecting lysosomal enzymes, transporters, or structural components, resulting in the accumulation of undegraded substrates and progressive cellular toxicity (173, 174). Among these, the neuronal ceroid lipofuscinoses (NCLs), caused by mutations in CLN genes such as CLN1 (also known as PPT1), CLN2 (also known as TPP1), CLN3, and CLN6 represent prototypical examples in which impaired lysosomal proteolysis and membrane trafficking lead to progressive neurodegeneration (175–179). Deficiency of GCase (GBA1) in Gaucher disease leads to accumulation of glucosylceramide and glucosylsphingosine, perturbing lysosomal membrane composition and trafficking (180–184).
In Niemann-Pick disease types A and B, loss of acid sphingomyelinase due to SMPD1 mutations causes sphingomyelin storage, whereas Niemann-Pick type C (NPC), resulting from NPC1 or NPC2 defects, impairs cholesterol and glycosphingolipid export, leading to unesterified cholesterol accumulation and defective endolysosomal motility (185, 186). Other LSDs, such as Tay-Sachs (HEXA mutation) and Fabry disease (GLA mutation), illustrate how deficits in ganglioside and glycosphingolipid degradation converge on lysosomal dysfunction and neurotoxicity (187).
Across these conditions, substrate overload distorts endolysosomal trafficking and acid hydrolase maturation, destabilizes LAMP2A, and thereby suppresses chaperone-mediated autophagy (188, 189). This disrupts lysosomal Ca²+ signaling through TRPML1, as seen in mucolipidosis type IV (190–192). Transcriptional responses driven by TFEB and TFE3 initially upregulate lysosomal biogenesis, but sustained activation may in certain contexts contribute to metabolic exhaustion and persistent inflammation, and its long-term impact remains under active investigation (193). Furthermore, recent evidence shows that STING acts as a lysosomal proton channel that directly activates TFEB under lysosomal stress, linking innate immune sensing to the transcriptional recovery of lysosomal function in storage disorders (194).
Thus, LSDs illustrate how impaired lipid homeostasis, defective repair, and failed autophagic adaptation produce a lysosomal fragility that mechanistically parallels the dysfunction seen in more common neurodegenerative diseases.
-
Organelle crosstalk, system integration, and future perspectives
Lysosomes exist within an integrated network of organelles that collectively sustain neuronal metabolism and stress resilience. Physical membrane contact sites connect lysosomes with other intracellular organelles, including the ER, mitochondria, peroxisomes, and the plasma membrane, creating a multidirectional circuit for lipid exchange, calcium signaling, and spatial positioning (195–198) (Figure 3). These dynamic contacts enable neurons to adapt rapidly to metabolic and mechanical stress, ensuring coordination with cellular energetic status. However, the molecular regulation of these contact sites and how their dynamics are altered during aging or disease remain incompletely characterized.
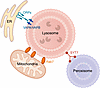 Figure 3
Figure 3Lysosomal inter-organelle contact sites. Lysosomes establish membrane contact sites with multiple organelles to coordinate metabolism and stress responses. At ER-lysosome interfaces, VAPA and VAPB engage ORPs to mediate cholesterol and phosphatidylserine exchange essential for membrane maintenance and repair. Mitochondria-lysosome contacts organized by Rab7 regulate calcium and lipid transfer, linking degradative capacity with mitochondrial energy supply. Peroxisome-lysosome junctions involving synaptotagmin 7 (SYT7) enable bidirectional lipid trafficking critical for plasmalogen and cholesterol homeostasis. Disruption of these connections impairs lipid balance and lysosomal positioning, sensitizing neurons to metabolic and oxidative stress.
At ER-lysosome interfaces, ER-resident VAPA and VAPB engage members of the ORP family, such as ORP9, ORP10, and ORP11, to mediate bidirectional transfer of cholesterol and PS in exchange for PI4P (48, 49) (Figure 3). This lipid flow maintains lysosomal membrane composition and provides precursors for the PI4K2A/ATG9A/ARFIP2 repair axis after injury (46, 49). During acute stress, ER tubules dynamically wrap around damaged lysosomes, establishing temporary bridges that deliver lipids and facilitate cholesterol-dependent resealing. Disruption of these tethers through the loss of VAPs or PI4K2A sensitizes cells to LMP, emphasizing the interdependence between lipid metabolism and membrane repair (48, 49).
Mitochondria-lysosome coupling is equally essential for maintaining cellular homeostasis (4, 196). Lysosomes remove dysfunctional mitochondria through mitophagy, while mitochondria supply ATP required for lysosomal acidification (199, 200). These organelles communicate through Rab7- and TBC1D15/17-dependent contact sites that coordinate calcium and lipid exchange (201) (Figure 3). Mitochondrial ROS can oxidize lysosomal membranes, whereas lysosomal rupture releases iron and cathepsins that promote mitochondrial permeabilization and apoptosis (202, 203). This bidirectional toxicity establishes a feed-forward loop that accelerates neuronal degeneration, particularly in PD and ALS (4).
Beyond these bilateral contacts, lysosomes also form tethers with peroxisomes, mediated by ORP1L and synaptotagmin VII (SYT7), enabling cholesterol and phospholipid exchange critical for maintaining plasmalogen and membrane lipid homeostasis (197) (Figure 3). Perturbation of this pathway contributes to the lipid dysregulation observed in neurodegenerative conditions. In addition, lysosome–plasma membrane contact sites regulate calcium-dependent exocytosis required for plasma membrane repair and clearance of aggregated proteins. This process involves SYT7, annexins, and SNARE family members such as vesicle-associated membrane protein 7 (VAMP7) and syntaxin 4 (198). Dysregulated lysosomal exocytosis contributes to the extracellular propagation of α-synuclein and tau aggregates in PD and AD (128).
Together, these diverse lysosomal contact sites, spanning the ER, mitochondria, peroxisomes, plasma membrane, and nucleus, form a spatially organized signaling network that integrates metabolism, membrane repair, and stress responses. Their disruption contributes to the defective lipid homeostasis, impaired clearance, and neuronal vulnerability that characterize multiple neurodegenerative diseases.
-
Aging and network decline
Aging leads to a gradual erosion of lysosomal quality control across multiple levels. The precise mechanisms driving this decline and whether they reflect intrinsic lysosomal aging or systemic metabolic changes remain open questions. The lipid composition of the lysosomal membrane shifts toward higher cholesterol and ceramide content, increasing rigidity and susceptibility to rupture (204, 205). Autophagic flux declines as the efficiency of autophagosome-lysosome fusion decreases, while TFEB activation becomes blunted due to persistent mTORC1 signaling and reduced calcineurin activity (206). These cumulative defects result in the buildup of lipofuscin, an autofluorescent aggregate composed of oxidized proteins and lipids that physically obstructs lysosomal degradation (207).
Mitochondrial output also wanes with age, reducing ATP availability for proton pumps and compromising lysosomal acidification (208). The number and stability of ER-lysosome and mitochondria-lysosome contact sites decline, limiting lipid and calcium transfer and weakening recovery from transient damage (209). This progressive uncoupling of organellar networks may underlie the early decline in neuronal proteostasis observed during aging, preceding overt neurodegeneration.
-
Therapeutic perspectives
Growing mechanistic insight into lysosomal homeostasis is reshaping therapeutic strategies for neurodegenerative diseases. Instead of focusing solely on downstream protein aggregates, emerging approaches aim to restore the lysosome’s intrinsic capacity to repair, replace, or regenerate itself (26, 210). However, although targeting lysosomal repair and regeneration pathways is conceptually promising, further work is needed to determine whether modulation of these mechanisms can alter disease trajectories in human patients.
Enhancing repair capacity represents one avenue. Small molecules that stabilize lysosomal membranes, modulate cholesterol distribution, or promote sphingomyelin–ceramide turnover have shown protective effects in models of neurodegeneration (211–213). Compounds that enhance ESCRT recruitment or boost lipid exchange through the PITT/ATG9A pathway may strengthen intrinsic repair mechanisms.
Reinforcing lysosomal clearance is another strategy. Pharmacological activation of TFEB or TFE3, via mTORC1 inhibition or calcineurin stimulation, upregulates genes that govern lysosomal biogenesis, autophagy, and lipid metabolism (214, 215). In experimental models of AD and PD, TFEB activation accelerates aggregate removal and improves neuronal viability (216, 217).
Targeting LRRK2 signaling holds promise for familial and sporadic PD. Selective LRRK2 kinase inhibitors normalize Rab phosphorylation, restore lysosomal trafficking, and are currently undergoing clinical testing (218, 219).
For LSDs and GBA1-associated PD, enzyme replacement and small-molecule chaperone therapies, such as ambroxol, improve substrate clearance and restore membrane stability (220, 221).
Finally, modulating lysosomal exocytosis may provide an adaptive outlet for cells under chronic stress. Controlled stimulation can aid in the removal of undegraded material and plasma membrane repair, although excessive activation risks propagating toxic aggregates and inflammatory signals.
Developing biomarkers of lysosomal health will be essential to monitor therapeutic efficacy and guide precision interventions.
-
Conclusion
Lysosomal homeostasis defines neuronal longevity by integrating stress sensing, repair, and regeneration into a unified survival program. While current evidence does not establish lysosomal repair failure as a universal initiating event in neurodegeneration, growing data support its role as a critical determinant of neuronal vulnerability and disease progression. Across neurodegenerative diseases, the failure of these adaptive mechanisms precipitates proteostatic collapse and inflammation. The convergence of ESCRT-mediated mechanics, lipid-based membrane repair, and TFEB-dependent transcriptional renewal reveals a common molecular architecture that underpins neuronal resilience. Restoring this triad through targeted modulation of lipid metabolism, membrane repair capacity, and transcriptional control offers a coherent strategy to halt neurodegeneration at its origin. Future work aimed at defining damage thresholds, pathway coordination, and neuron-specific vulnerabilities will be essential for translating these insights into therapeutic strategies.
-
Funding support
- Cancer Research UK grant CC2134 (to the Francis Crick Institute).
- UK Medical Research Council grant CC2034 (to the Francis Crick Institute).
- Wellcome Trust grant CC2134 (to the Francis Crick Institute).
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2026, De Tito et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: J Clin Invest. 2026;136(7):e199845. https://doi.org/10.1172/JCI199845.
-
References
- Scott O, et al. The spectrum of lysosomal stress and damage responses: from mechanosensing to inflammation. EMBO Rep. 2025;26(6):1425–1439.
- Settembre C, Perera RM. Lysosomes as coordinators of cellular catabolism, metabolic signalling and organ physiology. Nat Rev Mol Cell Biol. 2024;25(3):223–245.
- Rizzollo F, et al. The lysosome as a master regulator of iron metabolism. Trends Biochem Sci. 2021;46(12):960–975.
- Deus CM, et al. Mitochondria-lysosome crosstalk: from physiology to neurodegeneration. Trends Mol Med. 2020;26(1):71–88.
- Ebner M, et al. Nutrient-regulated control of lysosome function by signaling lipid conversion. Cell. 2023;186(24):5328–5346.
- Radulovic M, et al. Lysosomal membrane homeostasis and its importance in physiology and disease. Nat Rev Mol Cell Biol. 2026;27(1):71–87.
- Napolitano G, Ballabio A. TFEB at a glance. J Cell Sci. 2016;129(13):2475–2481.
- Carroll B, Dunlop EA. The lysosome: a crucial hub for AMPK and mTORC1 signalling. Biochem J. 2017;474(9):1453–1466.
- Karpova A, et al. Neuronal autophagy in the control of synapse function. Neuron. 2025;113(7):974–990.
- Nixon RA. Autophagy-lysosomal-associated neuronal death in neurodegenerative disease. Acta Neuropathol. 2024;148(1):42.
- Lie PPY, Nixon RA. Lysosome trafficking and signaling in health and neurodegenerative diseases. Neurobiol Dis. 2019;122:94–105.
- Konietzny A, et al. Efficient axonal transport of endolysosomes relies on the balanced ratio of microtubule tyrosination and detyrosination. J Cell Sci. 2024;137(8):jcs261737.
- Lee S, et al. Lysosomal proteolysis inhibition selectively disrupts axonal transport of degradative organelles and causes an Alzheimer’s-like axonal dystrophy. J Neurosci. 2011;31(21):7817–7830.
- Udayar V, et al. Lysosomal dysfunction in neurodegeneration: emerging concepts and methods. Trends Neurosci. 2022;45(3):184–199.
- Dai L, et al. Lysosomal dysfunction in α-synuclein pathology: molecular mechanisms and therapeutic strategies. Cell Mol Life Sci. 2024;81(1):382.
- Nguyen M, et al. Synaptic, mitochondrial, and lysosomal dysfunction in Parkinson’s disease. Trends Neurosci. 2019;42(2):140–149.
- Root J, et al. Lysosome dysfunction as a cause of neurodegenerative diseases: Lessons from frontotemporal dementia and amyotrophic lateral sclerosis. Neurobiol Dis. 2021;154:105360.
- Krogsaeter EK, et al. Lysosomal proteomics reveals mechanisms of neuronal APOE4-associated lysosomal dysfunction. Autophagy. 2025;21(12):3240–3265.
- Sampognaro PJ, et al. Mutations in α-synuclein, TDP-43 and tau prolong protein half-life through diminished degradation by lysosomal proteases. Mol Neurodegener. 2023;18(1):29.
- Ratto E, et al. Direct control of lysosomal catabolic activity by mTORC1 through regulation of V-ATPase assembly. Nat Commun. 2022;13(1):4848.
- Saimoto Y, et al. Lysosomal lipid peroxidation contributes to ferroptosis induction via lysosomal membrane permeabilization. Nat Commun. 2025;16(1):3554.
- He X, et al. The role of redox-mediated lysosomal dysfunction and therapeutic strategies. Biomed Pharmacother. 2023;165:115121.
- Zoncu R, Perera RM. Built to last: lysosome remodeling and repair in health and disease. Trends Cell Biol. 2022;32(7):597–610.
- Lei Y, Klionsky DJ. A delicate decision between repair and degradation of damaged lysosomes. Autophagy. 2024;20(7):1471–1472.
- Otomo T, Yoshimori T. Lysophagy: a method for monitoring lysosomal rupture followed by autophagy-dependent recovery. Methods Mol Biol. 2017;1594:141–149.
- Xun J, Tan JX. Lysosomal repair in health and disease. J Cell Physiol. 2025;240(5):e70044.
- Hari A, et al. Activation of NLRP3 inflammasome by crystalline structures via cell surface contact. Sci Rep. 2014;4(1):7281.
- Bussi C, et al. Lysosomal damage drives mitochondrial proteome remodelling and reprograms macrophage immunometabolism. Nat Commun. 2022;13(1):7338.
- Wang F, et al. Lysosomal membrane permeabilization and cell death. Traffic. 2018;19(12):918–931.
- Wang F, et al. Lysosomal membrane permeabilization and cell death. Traffic Cph Den. 2018;19(12):918–931.
- Yamashima T. Lysosomal membrane-permeabilization (LMP) and -rupture (LMR) are distinct for cell death. Front Cell Death. 2025;4:1669955.
- Jia J, et al. Galectins control mTOR in response to endomembrane damage. Mol Cell. 2018;70(1):120–135.
- Jia J, et al. Galectin-3 coordinates a cellular system for lysosomal repair and removal. Dev Cell. 2020;52(1):69–87.
- Shin HR, Zoncu R. Finding sugar in the pantry: how galectins detect and signal lysosomal damage. Mol Cell. 2018;70(1):5–7.
- Sudhakar JN, et al. Lumenal Galectin-9-Lamp2 interaction regulates lysosome and autophagy to prevent pathogenesis in the intestine and pancreas. Nat Commun. 2020;11(1):4286.
- Radulovic M, et al. ESCRT-mediated lysosome repair precedes lysophagy and promotes cell survival. EMBO J. 2018;37(21):e99753.
- Duran J, et al. Calcium signaling from damaged lysosomes induces cytoprotective stress granules. EMBO J. 2024;43(24):6410–6443.
- Chen W, et al. Ca2+-sensor ALG-2 engages ESCRTs to enhance lysosomal membrane resilience to osmotic stress. Proc Natl Acad Sci U S A. 2024;121(22):e2318412121. View this article via: CrossRef Google Scholar
- Sun S, et al. ALG-2 activates the MVB sorting function of ALIX through relieving its intramolecular interaction. Cell Discov. 2015;1(1):15018.
- Stuchell-Brereton MD, et al. ESCRT-III recognition by VPS4 ATPases. Nature. 2007;449(7163):740–744.
- Han H, et al. The AAA ATPase Vps4 binds ESCRT-III substrates through a repeating array of dipeptide-binding pockets. Elife. 2017;6:e31324.
- Quick JD, et al. Lysosomal acidification dysfunction in microglia: an emerging pathogenic mechanism of neuroinflammation and neurodegeneration. J Neuroinflammation. 2023;20(1):185.
- Chou C-C, et al. Proteostasis and lysosomal repair deficits in transdifferentiated neurons of Alzheimer’s disease. Nat Cell Biol. 2025;27(4):619–632.
- Jia J, et al. AMPK, a regulator of metabolism and autophagy, is activated by lysosomal damage via a novel galectin-directed ubiquitin signal transduction system. Mol Cell. 2020;77(5):951–969.
- Jia J, et al. AMPK is activated during lysosomal damage via a galectin-ubiquitin signal transduction system. Autophagy. 2020;16(8):1550–1552.
- De Tito S, et al. ATG9A and ARFIP2 cooperate to control PI4P levels for lysosomal repair. Dev Cell. 2025;60(20):2744–2760.
- Meyer H, Kravic B. ATG9 assists lipid replenishment for lysosome membrane repair. Dev Cell. 2025;60(20):2701–2702.
- Goul CS, Zoncu R. PITTching in for lysosome repair. Dev Cell. 2022;57(20):2347–2349.
- Tan JX, Finkel T. A phosphoinositide signalling pathway mediates rapid lysosomal repair. Nature. 2022;609(7928):815–821.
- Peng K, et al. The autophagy protein ATG-9 regulates lysosome function and integrity. J Cell Biol. 2025;224(6):e202411092.
- Radulovic M, et al. Cholesterol transfer via endoplasmic reticulum contacts mediates lysosome damage repair. EMBO J. 2022;41(24):e112677.
- Sawa-Makarska J, et al. Reconstitution of autophagosome nucleation defines Atg9 vesicles as seeds for membrane formation. Science. 2020;369(6508):eaaz7714.
- Guardia CM, et al. Structure of human ATG9A, the only transmembrane protein of the core autophagy machinery. Cell Rep. 2020;31(13):107837.
- Millard E, Tooze SA. ATG9 not just an autophagy related protein. J Mol Biol. 2025;437(18):169288.
- Vliet AR van, et al. ATG9A and ATG2A form a heteromeric complex essential for autophagosome formation. Mol Cell. 2022;82(22):4324–4339.
- De Tito S, et al. The Golgi as an assembly line to the autophagosome. Trends Biochem Sci. 2020;45(6):484–496.
- Claude-Taupin A, et al. ATG9A protects the plasma membrane from programmed and incidental permeabilization. Nat Cell Biol. 2021;23(8):846–858.
- Ambroggio EE, et al. Arf1 and membrane curvature cooperate to recruit Arfaptin2 to liposomes. PLoS One. 2013;8(4):e62963.
- Nakamura K, et al. Structural basis for membrane binding specificity of the Bin/Amphiphysin/Rvs (BAR) domain of Arfaptin-2 determined by Arl1 GTPase. J Biol Chem. 2012;287(30):25478–25489.
- Cruz-Garcia D, et al. Recruitment of arfaptins to the trans-Golgi network by PI(4)P and their involvement in cargo export. EMBO J. 2013;32(12):1717–1729.
- Rocha EM, et al. LRRK2 inhibition prevents endolysosomal deficits seen in human Parkinson’s disease. Neurobiol Dis. 2020;134:104626.
- Erb ML, Moore DJ. LRRK2 and the endolysosomal system in parkinson’s disease. J Park Dis. 2020;10(4):1271–1291.View this article via: PubMed Google Scholar
- Twellsieck B, Boecker CA. Role of LRRK2 in axonal transport and Parkinson’s disease. Biochem J. 2025;482(13):905–919.
- Niekamp P, et al. Ca2+-activated sphingomyelin scrambling and turnover mediate ESCRT-independent lysosomal repair. Nat Commun. 2022;13(1):1875.
- Varela YR, et al. Ceramide and the membrane-fusion activity of LC3/GABARAP autophagy proteins. Cell Mol Life Sci. 2025;82(1):283.
- Du X, et al. A role for oxysterol-binding protein-related protein 5 in endosomal cholesterol trafficking. J Cell Biol. 2011;192(1):121–135.
- Bussi C, et al. Stress granules plug and stabilize damaged endolysosomal membranes. Nature. 2023;623(7989):1062–1069.
- Bonet-Ponce L, et al. Mechanisms of lysosomal tubulation and sorting driven by LRRK2. Biochem Soc Trans. 2024;52(4):1909–1919.
- Melia TJ, et al. Autophagosome biogenesis: from membrane growth to closure. J Cell Biol. 2020;219(6):e202002085.
- Axe EL, et al. Autophagosome formation from membrane compartments enriched in phosphatidylinositol 3-phosphate and dynamically connected to the endoplasmic reticulum. J Cell Biol. 2008;182(4):685–701.
- Gómez-Sánchez R, et al. Establishment of the phagophore-ERES membrane contact site initiates phagophore elongation. Nat Struct Mol Biol. 2025;32(11):2319–2334.
- Nishimura T, Tooze SA. Emerging roles of ATG proteins and membrane lipids in autophagosome formation. Cell Discov. 2020;6(1):32.
- Grumati P, Dikic I. Ubiquitin signaling and autophagy. J Biol Chem. 2018;293(15):5404–5413.
- Gubas A, Dikic I. A guide to the regulation of selective autophagy receptors. FEBS J. 2022;289(1):75–89.
- Eapen VV, et al. Quantitative proteomics reveals the selectivity of ubiquitin-binding autophagy receptors in the turnover of damaged lysosomes by lysophagy. Elife. 2021;10:e72328.
- Gahlot P, et al. Lysosomal damage sensing and lysophagy initiation by SPG20-ITCH. Mol Cell. 2024;84(8):1556–1569.
- Park NY, et al. Activation of lysophagy by a TBK1-SCFFBXO3-TMEM192-TAX1BP1 axis in response to lysosomal damage. Nat Commun. 2025;16(1):1109.
- Koerver L, et al. The ubiquitin-conjugating enzyme UBE2QL1 coordinates lysophagy in response to endolysosomal damage. EMBO Rep. 2019;20(10):e48014.
- Mizushima N. The ubiquitin E2 enzyme UBE2QL1 mediates lysophagy. EMBO Rep. 2019;20(10):e49104.
- Yoshida Y, et al. Ubiquitination of exposed glycoproteins by SCFFBXO27 directs damaged lysosomes for autophagy. Proc Natl Acad Sci U S A. 2017;114(32):8574–8579.
- Teranishi H, et al. Identification of CUL4A-DDB1-WDFY1 as an E3 ubiquitin ligase complex involved in initiation of lysophagy. Cell Rep. 2022;40(11):111349.
- Chauhan S, et al. TRIMs and galectins globally cooperate and TRIM16 and galectin-3 co-direct autophagy in endomembrane damage homeostasis. Dev Cell. 2016;39(1):13–27.
- Thurston TLM, et al. Galectin 8 targets damaged vesicles for autophagy to defend cells against bacterial invasion. Nature. 2012;482(7385):414–418.
- Boyle KB, et al. CALCOCO2/NDP52 initiates selective autophagy through recruitment of ULK and TBK1 kinase complexes. Autophagy. 2019;15(9):1655–1656.
- Gallagher ER, Holzbaur ELF. The selective autophagy adaptor p62/SQSTM1 forms phase condensates regulated by HSP27 that facilitate the clearance of damaged lysosomes via lysophagy. Cell Rep. 2023;42(2):112037.
- Bussi C, et al. Alpha-synuclein fibrils recruit TBK1 and OPTN to lysosomal damage sites and induce autophagy in microglial cells. J Cell Sci. 2018;131(23):jcs226241.
- Fujita N, et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol Biol Cell. 2008;19(5):2092–2100.
- Kravić B, et al. Ubiquitin profiling of lysophagy identifies actin stabilizer CNN2 as a target of VCP/p97 and uncovers a link to HSPB1. Mol Cell. 2022;82(14):2633–2649.
- Medina DL, et al. Lysosomal calcium signalling regulates autophagy through calcineurin and TFEB. Nat Cell Biol. 2015;17(3):288–299.
- Xia Q, et al. SMURF1 controls the PPP3/calcineurin complex and TFEB at a regulatory node for lysosomal biogenesis. Autophagy. 20(4):735–751.
- Meireles AM, et al. The lysosomal transcription factor TFEB represses myelination downstream of the rag-ragulator complex. Dev Cell. 2018;47(3):319–330.
- Zou W, et al. Lysosomal dynamics regulate mammalian cortical neurogenesis. Dev Cell. 2024;59(1):64–78.
- Zhang Y, et al. The cGAS-STING pathway drives neuroinflammation and neurodegeneration via cellular and molecular mechanisms in neurodegenerative diseases. Neurobiol Dis. 2024;202:106710.
- Mächtel R, et al. From lysosomal storage disorders to Parkinson’s disease - challenges and opportunities. J Mol Biol. 2023;435(12):167932.
- Rusmini P, et al. Impairment of lysosomal quality control in Huntington disease. Cell Death Dis. 2025;16(1):762.
- Hampel H, et al. The amyloid-β pathway in Alzheimer’s Disease. Mol Psychiatry. 2021;26(10):5481–5503.
- Nixon RA. Amyloid precursor protein and endosomal-lysosomal dysfunction in Alzheimer’s disease: inseparable partners in a multifactorial disease. FASEB J. 2017;31(7):2729–2743.
- Arafi P, et al. Alzheimer-mutant γ-secretase complexes stall amyloid β-peptide production. Elife. 2025;13:RP102274.
- Deaton CA, Johnson GVW. Presenilin 1 regulates membrane homeostatic pathways that are dysregulated in Alzheimer’s disease. J Alzheimers Dis. 2020;77(3):961–977.
- Fedeli C, et al. PSEN2 (presenilin 2) mutants linked to familial Alzheimer disease impair autophagy by altering Ca2+ homeostasis. Autophagy. 2019;15(12):2044–2062.
- Meckler X, Checler F. Presenilin 1 and presenilin 2 target γ-secretase complexes to distinct cellular compartments. J Biol Chem. 2016;291(24):12821–12837.
- Lee JH, et al. Lysosomal proteolysis and autophagy require presenilin 1 and are disrupted by Alzheimer-related PS1 mutations. Cell. 2010;141(7):1146–1158.
- Cai Y, et al. Mutations in presenilin 2 and its implications in Alzheimer’s disease and other dementia-associated disorders. Clin Interv Aging. 2015;10:1163–1172.View this article via: PubMed Google Scholar
- Delabio R, et al. PSEN1 and PSEN2 gene expression in Alzheimer’s disease brain: a new approach. J Alzheimers Dis. 2014;42(3):757–760.
- Xia D, et al. Presenilin-1 knockin mice reveal loss-of-function mechanism for familial Alzheimer’s disease. Neuron. 2015;85(5):967–981.
- Chen X, et al. Caffeine blocks disruption of blood brain barrier in a rabbit model of Alzheimer’s disease. J Neuroinflammation. 2008;5:12.
- Takechi R, et al. Three-dimensional colocalization analysis of plasma-derived apolipoprotein B with amyloid plaques in APP/PS1 transgenic mice. Histochem Cell Biol. 2009;131(5):661–666.
- Chen X, et al. Endolysosome mechanisms associated with Alzheimer’s disease-like pathology in rabbits ingesting cholesterol-enriched diet. J Alzheimers Dis. 2010;22(4):1289–1303.
- Hui L, et al. Endolysosome involvement in LDL cholesterol-induced Alzheimer’s disease-like pathology in primary cultured neurons. Life Sci. 2012;91(23-24):1159–1168.
- Hellén M, et al. Inflammation-induced lysosomal dysfunction in human iPSC-derived microglia is exacerbated by APOE 4/4 genotype. J Neuroinflammation. 2025;22(1):147.
- Litvinchuk A, et al. Amelioration of Tau and ApoE4-linked glial lipid accumulation and neurodegeneration with an LXR agonist. Neuron. 2024;112(12):2079.
- Connolly KJ, et al. Loss of lipid carrier ApoE exacerbates brain glial and inflammatory responses after lysosomal GBA1 inhibition. Cells. 2023;12(21):2564.
- Nixon RA. Autophagy, amyloidogenesis and Alzheimer disease. J Cell Sci. 2007;120(pt 23):4081–4091.
- Nixon RA, et al. Extensive involvement of autophagy in Alzheimer disease: an immuno-electron microscopy study. J Neuropathol Exp Neurol. 2005;64(2):113–122.
- Rose K, et al. Tau fibrils induce nanoscale membrane damage and nucleate cytosolic tau at lysosomes. Proc Natl Acad Sci U S A. 2024;121(22):e2315690121.
- Li T, et al. TFEB acetylation promotes lysosome biogenesis and ameliorates Alzheimer’s disease–relevant phenotypes in mice. J Biol Chem. 2022;298(12):102649.
- Gu Z, et al. TFEB in Alzheimer’s disease: From molecular mechanisms to therapeutic implications. Neurobiol Dis. 2022;173:105855.
- Bretou M, et al. Accumulation of APP C-terminal fragments causes endolysosomal dysfunction through the dysregulation of late endosome to lysosome-ER contact sites. Dev Cell. 2024;59(12):1571–1592.
- Filippov V, et al. Increased ceramide in brains with Alzheimer’s and other neurodegenerative diseases. J Alzheimers Dis. 2012;29(3):537–547.
- Vidyadhara DJ, et al. Role of the endolysosomal system in Parkinson’s disease. J Neurochem. 2019;150(5):487–506.
- Chatterjee D, Krainc D. Mechanisms of glucocerebrosidase dysfunction in Parkinson’s disease. J Mol Biol. 2023;435(12):168023.
- Farfel-Becker T, et al. Can GBA1-associated Parkinson disease be modeled in the mouse? Trends Neurosci. 2019;42(9):631–643.
- Collins LM, et al. Dermal fibroblasts from patients with Parkinson’s disease have normal GCase activity and autophagy compared to patients with PD and GBA mutations. F1000Res. 2018;6:1751.
- Tang Q, et al. Alpha-Synuclein defects autophagy by impairing SNAP29-mediated autophagosome-lysosome fusion. Cell Death Dis. 2021;12(10):854.
- Flavin WP, et al. Endocytic vesicle rupture is a conserved mechanism of cellular invasion by amyloid proteins. Acta Neuropathol. 2017;134(4):629–653.
- Ho MS. Clearance pathways for α-synuclein in Parkinson’s disease. J Neurochem. 2025;169(6):e70124.
- Xie YX, et al. Lysosomal exocytosis releases pathogenic α-synuclein species from neurons in synucleinopathy models. Nat Commun. 2022;13(1):4918.
- Samaddar M, et al. Lysosomal polyamine storage upon ATP13A2 loss impairs β-glucocerebrosidase via altered lysosomal pH and electrostatic hydrolase-lipid interactions. Cell Rep. 2025;44(9):116179.
- Dehay B, et al. Loss of P-type ATPase ATP13A2/PARK9 function induces general lysosomal deficiency and leads to Parkinson disease neurodegeneration. Proc Natl Acad Sci U S A. 2012;109(24):9611–9616.
- Zhang F, et al. The Roles of ATP13A2 gene mutations leading to abnormal aggregation of α-synuclein in Parkinson’s disease. Front Cell Neurosci. 2022;16:927682.
- Hu M, et al. Parkinson’s disease-risk protein TMEM175 is a proton-activated proton channel in lysosomes. Cell. 2022;185(13):2292–2308.
- Jinn S, et al. TMEM175 deficiency impairs lysosomal and mitochondrial function and increases α-synuclein aggregation. Proc Natl Acad Sci U S A. 2017;114(9):2389–2394.
- Wu L, et al. TMEM175: A lysosomal ion channel associated with neurological diseases. Neurobiol Dis. 2023;185:106244.
- Rowlands J, Moore DJ. VPS35 and retromer dysfunction in Parkinson’s disease. Philos Trans R Soc Lond B Biol Sci. 2024;379(1899):20220384.
- Tsika E, et al. Parkinson’s disease-linked mutations in VPS35 induce dopaminergic neurodegeneration. Hum Mol Genet. 2014;23(17):4621–4638.
- Zavodszky E, et al. Mutation in VPS35 associated with Parkinson’s disease impairs WASH complex association and inhibits autophagy. Nat Commun. 2014;5(1):3828.
- Su J, et al. Systemic knockout of Tmem175 results in aberrant differentiation but no effect on hematopoietic reconstitution. Stem Cell Res. 2024;79:103469.
- Todd TW, et al. The endolysosomal pathway and ALS/FTD. Trends Neurosci. 2023;46(12):1025–1041.
- Hasan S, et al. Multi-modal proteomic characterization of lysosomal function and proteostasis in progranulin-deficient neurons. Mol Neurodegener. 2023;18(1):87.
- Simon MJ, et al. Lysosomal functions of progranulin and implications for treatment of frontotemporal dementia. Trends Cell Biol. 2023;33(4):324–339.
- Tanaka Y, et al. Progranulin regulates lysosomal function and biogenesis through acidification of lysosomes. Hum Mol Genet. 2017;26(5):969–988.View this article via: PubMed Google Scholar
- Goodier JL, et al. C9orf72-associated SMCR8 protein binds in the ubiquitin pathway and with proteins linked with neurological disease. Acta Neuropathol Commun. 2020;8(1):110.
- Dong D, et al. Poly-GR repeats associated with ALS/FTD gene C9ORF72 impair translation elongation and induce a ribotoxic stress response in neurons. Sci Signal. 2024;17(848):eadl1030.
- Frottin F, et al. Multiple pathways of toxicity induced by C9orf72 dipeptide repeat aggregates and G4C2 RNA in a cellular model. Elife. 2021;10:e62718.
- Lee Y-B, et al. C9orf72 poly GA RAN-translated protein plays a key role in amyotrophic lateral sclerosis via aggregation and toxicity. Hum Mol Genet. 2017;26(24):4765–4777.
- Marchi PM, et al. C9ORF72-derived poly-GA DPRs undergo endocytic uptake in iAstrocytes and spread to motor neurons. Life Sci Alliance. 2022;5(9):e202101276.
- Ferrari V, et al. Pathogenic variants of Valosin-containing protein induce lysosomal damage and transcriptional activation of autophagy regulators in neuronal cells. Neuropathol Appl Neurobiol. 2022;48(5):e12818.
- Clayton EL, et al. Frontotemporal dementia caused by CHMP2B mutation is characterised by neuronal lysosomal storage pathology. Acta Neuropathol. 2015;130(4):511–523.
- van der Zee J, et al. CHMP2B C-truncating mutations in frontotemporal lobar degeneration are associated with an aberrant endosomal phenotype in vitro. Hum Mol Genet. 2008;17(2):313–322.
- Ferrari R, et al. Novel missense mutation in charged multivesicular body protein 2B in a patient with frontotemporal dementia. Alzheimer Dis Assoc Disord. 2010;24(4):397–401.View this article via: CrossRef Google Scholar
- Harding O, et al. ALS- and FTD-associated missense mutations in TBK1 differentially disrupt mitophagy. Proc Natl Acad Sci U S A. 2021;118(24):e2025053118.
- Ye J, et al. Effects of ALS-associated TANK binding kinase 1 mutations on protein-protein interactions and kinase activity. Proc Natl Acad Sci U S A. 2019;116(49):24517–24526.
- Gerbino V, et al. The loss of TBK1 kinase activity in motor neurons or in all cell types differentially impacts ALS disease progression in SOD1 mice. Neuron. 2020;106(5):789–805.
- Ji YJ, et al. C9orf72/ALFA-1 controls TFEB/HLH-30-dependent metabolism through dynamic regulation of Rag GTPases. PLoS Genet. 2020;16(4):e1008738.
- Borrego-Écija S, et al. Galectin-3 is upregulated in frontotemporal dementia patients with subtype specificity. Alzheimers Dement. 2024;20(3):1515–1526.
- Jiang A, et al. From pathogenesis to therapeutics: a review of 150 years of Huntington’s disease research. Int J Mol Sci. 2023;24(16):13021.
- Tabrizi SJ, et al. Huntington disease: new insights into molecular pathogenesis and therapeutic opportunities. Nat Rev Neurol. 2020;16(10):529–546.
- Rui Y-N, et al. HTT/Huntingtin in selective autophagy and Huntington disease: A foe or a friend within? Autophagy. 2015;11(5):858–860.
- Krzystek TJ, et al. HTT (huntingtin) and RAB7 co-migrate retrogradely on a signaling LAMP1-containing late endosome during axonal injury. Autophagy. 2023;19(4):1199–1220.
- Caviston JP, et al. Huntingtin coordinates the dynein-mediated dynamic positioning of endosomes and lysosomes. Mol Biol Cell. 2011;22(4):478–492.
- Martinez-Vicente M, et al. Cargo recognition failure is responsible for inefficient autophagy in Huntington’s disease. Nat Neurosci. 2010;13(5):567–576.
- Wong YC, Holzbaur ELF. The regulation of autophagosome dynamics by huntingtin and HAP1 is disrupted by expression of mutant huntingtin, leading to defective cargo degradation. J Neurosci. 2014;34(4):1293–1305.
- Pryor WM, et al. Huntingtin promotes mTORC1 signaling in the pathogenesis of Huntington’s disease. Sci Signal. 2014;7(349):ra103.
- Stavrides P, et al. mTOR inhibition in Q175 Huntington’s disease model mice facilitates neuronal autophagy and mutant huntingtin clearance. Elife. 2025;14:RP104979.
- Kacher R, et al. Altered cholesterol homeostasis in Huntington’s disease. Front Aging Neurosci. 2022;14:797220.
- Phillips GR, et al. Phospholipid profiles are selectively altered in the putamen and white frontal cortex of Huntington’s disease. Nutrients. 2022;14(10):2086.
- Fraldi A, et al. Lysosomal fusion and SNARE function are impaired by cholesterol accumulation in lysosomal storage disorders. EMBO J. 2010;29(21):3607–3620.
- Sameni S, et al. Alteration in fluidity of cell plasma membrane in Huntington disease revealed by spectral phasor analysis. Sci Rep. 2018;8(1):734.
- Di Pardo A, et al. De novo synthesis of sphingolipids is defective in experimental models of Huntington’s disease. Front Neurosci. 2017;11:698.
- Pal A, et al. Huntingtin-HAP40 complex is a novel Rab5 effector that regulates early endosome motility and is up-regulated in Huntington’s disease. J Cell Biol. 2006;172(4):605–618.
- Yang J, et al. A prion-like domain of TFEB mediates the co-aggregation of TFEB and mHTT. Autophagy. 2023;19(2):544–550.
- Siew JJ, et al. Galectin-3 is required for the microglia-mediated brain inflammation in a model of Huntington’s disease. Nat Commun. 2019;10(1):3473.
- Sheth J, et al Lysosomal storage disorders: from biology to the clinic with reference to India. Lancet Reg Health Southeast Asia. 2023;9:100108. View this article via: PubMed Google Scholar
- Platt FM, et al. Lysosomal storage diseases. Nat Rev Dis Primer. 2018;4(1):27.
- Alam MS, et al. AAV-delivered PPT1 provides long-term neurological benefits in CLN1 mice and achieves therapeutic levels in sheep brain. Mol Ther. 2025;33(10):4799–4819.
- Geraets RD, et al. A tailored mouse model of CLN2 disease: a nonsense mutant for testing personalized therapies. PLoS One. 2017;12(5):e0176526.
- Gardner E, et al. Mutation update: review of TPP1 gene variants associated with neuronal ceroid lipofuscinosis CLN2 disease. Hum Mutat. 2019;40(11):1924–1938.
- Singh JB, et al. CLN3 disease disrupts very early postnatal hippocampal maturation. Sci Rep. 2025;15(1):24411.
- Cain JT, et al. Gene therapy corrects brain and behavioral pathologies in CLN6-batten disease. Mol Ther. 2019;27(10):1836–1847.
- Mistry PK, et al. Glucocerebrosidase gene-deficient mouse recapitulates Gaucher disease displaying cellular and molecular dysregulation beyond the macrophage. Proc Natl Acad Sci U S A. 2010;107(45):19473–19478.
- Marano M, et al. Increased glucosylsphingosine levels and Gaucher disease in GBA1-associated Parkinson’s disease. Parkinsonism Relat Disord. 2024;124:107023.
- Pitcairn C, et al. Dysregulation of the autophagic-lysosomal pathway in Gaucher and Parkinson’s disease. Neurobiol Dis. 2019;122:72–82.
- Xu YH, et al. Viable mouse models of acid beta-glucosidase deficiency: the defect in Gaucher disease. Am J Pathol. 2003;163(5):2093–2101.
- Wen S, et al. A novel mouse model of chronic neuronopathic Gaucher disease exhibits Parkinson’s disease-like phenotypes. Neurobiol Dis. 2025;209:106899.
- McCauliff LA, et al. Intracellular cholesterol trafficking is dependent upon NPC2 interaction with lysobisphosphatidic acid. Elife. 2019;8:e50832.
- Li J, et al. Glycosylation inhibition reduces cholesterol accumulation in NPC1 protein-deficient cells. Proc Natl Acad Sci U S A. 2015;112(48):14876–14881.
- Xu YH, et al. Multi-system disorders of glycosphingolipid and ganglioside metabolism. J Lipid Res. 2010;51(7):1643–1675.
- Li W, et al. Phosphorylation of LAMP2A by p38 MAPK couples ER stress to chaperone-mediated autophagy. Nat Commun. 2017;8(1):1763.
- Grochowska KM, et al. Chaperone-mediated autophagy in neuronal dendrites utilizes activity-dependent lysosomal exocytosis for protein disposal. Cell Rep. 2023;42(8):112998.
- Cao Q, et al. The lysosomal Ca2+ release channel TRPML1 regulates lysosome size by activating calmodulin. J Biol Chem. 2017;292(20):8424–8435.
- Dong XP, et al. The type IV mucolipidosis-associated protein TRPML1 is an endolysosomal iron release channel. Nature. 2008;455(7215):992–996.
- Micsenyi MC, et al. Neuropathology of the Mcoln1(-/-) knockout mouse model of mucolipidosis type IV. J Neuropathol Exp Neurol. 2009;68(2):125–135.
- Chen H, et al. From the regulatory mechanism of TFEB to its therapeutic implications. Cell Death Discov. 2024;10(1):84.
- Tang Z, et al. STING mediates lysosomal quality control and recovery through its proton channel function and TFEB activation in lysosomal storage disorders. Mol Cell. 2025;85(8):1624–1639.
- Bandyopadhyay S, et al. Lysosomal membrane contact sites: Integrative hubs for cellular communication and homeostasis. Curr Top Membr. 2024;93:85–116.
- Cisneros J, et al. Mitochondria-lysosome contact site dynamics and misregulation in neurodegenerative diseases. Trends Neurosci. 2022;45(4):312–322.
- Chu B-B, et al. Cholesterol transport through lysosome-peroxisome membrane contacts. Cell. 2015;161(2):291–306.
- Rao SK, et al. Identification of SNAREs involved in synaptotagmin VII-regulated lysosomal exocytosis. J Biol Chem. 2004;279(19):20471–20479.
- Evans CS, Holzbaur ELF. Lysosomal degradation of depolarized mitochondria is rate-limiting in OPTN-dependent neuronal mitophagy. Autophagy. 2020;16(5):962–964.
- Yagi M, et al. Mitochondrial translation deficiency impairs NAD+ -mediated lysosomal acidification. EMBO J. 2021;40(8):e105268.
- Jongsma ML, et al. SKIP-HOPS recruits TBC1D15 for a Rab7-to-Arl8b identity switch to control late endosome transport. EMBO J. 2020;39(6):e102301.
- Oberle C, et al. Lysosomal membrane permeabilization and cathepsin release is a Bax/Bak-dependent, amplifying event of apoptosis in fibroblasts and monocytes. Cell Death Differ. 2010;17(7):1167–1178.
- Ghosh M, et al. Lysosomal membrane permeabilization causes oxidative stress and ferritin induction in macrophages. FEBS Lett. 2011;585(4):623–629.
- Nixon RA. The aging lysosome: An essential catalyst for late-onset neurodegenerative diseases. Biochim Biophys Acta Proteins Proteom. 2020;1868(9):140443.
- Tan JX, Finkel T. Lysosomes in senescence and aging. EMBO Rep. 2023;24(11):e57265.
- Aman Y, et al. Autophagy in healthy aging and disease. Nat Aging. 2021;1(8):634–650.
- Pan C, et al. Lipofuscin causes atypical necroptosis through lysosomal membrane permeabilization. Proc Natl Acad Sci U S A. 2021;118(47):e2100122118.
- Somasundaram I, et al. Mitochondrial dysfunction and its association with age-related disorders. Front Physiol. 2024;15:1384966.
- Azarnia Tehran D, Pizzo P. The complex web of membrane contact sites in brain aging and neurodegeneration. Cell Mol Life Sci. 2025;82(1):301.
- Bonam SR, et al. Lysosomes as a therapeutic target. Nat Rev Drug Discov. 2019;18(12):923–948.
- Hippman RS, et al. Discovery of a small-molecule modulator of the autophagy-lysosome pathway that targets lamin A/C and LAMP1, induces autophagic flux, and affects lysosome positioning in neurons. ACS Chem Neurosci. 2023;14(24):4363–4382.
- Baloni P, et al. Multi-omic analyses characterize the ceramide/sphingomyelin pathway as a therapeutic target in Alzheimer’s disease. Commun Biol. 2022;5(1):1074.
- Liu B, et al. Cyclodextrin overcomes the transport defect in nearly every organ of NPC1 mice leading to excretion of sequestered cholesterol as bile acid. J Lipid Res. 2010;51(5):933–944.
- Zhitomirsky B, et al. Lysosomotropic drugs activate TFEB via lysosomal membrane fluidization and consequent inhibition of mTORC1 activity. Cell Death Dis. 2018;9(12):1191.
- Du K, et al. Small-molecule activation of TFEB alleviates Niemann-Pick disease type C via promoting lysosomal exocytosis and biogenesis. Elife. 2025;13:RP103137.
- Song JX, et al. A small molecule transcription factor EB activator ameliorates beta-amyloid precursor protein and Tau pathology in Alzheimer’s disease models. Aging Cell. 2020;19(2):e13069.
- Lin J, et al. TFEB agonist clomiphene citrate activates the autophagy-lysosomal pathway and ameliorates Alzheimer’s disease symptoms in mice. J Biol Chem. 2024;300(12):107929.
- Jennings D, et al. Preclinical and clinical evaluation of the LRRK2 inhibitor DNL201 for Parkinson’s disease. Sci Transl Med. 2022;14(648):eabj2658.
- Zhu H, et al. Pharmacology of LRRK2 with type I and II kinase inhibitors revealed by cryo-EM. Cell Discov. 2024;10(1):10.
- Mullin S, et al. Ambroxol for the treatment of patients with Parkinson disease with and without glucocerebrosidase gene mutations: a nonrandomized, noncontrolled trial. JAMA Neurol. 2020;77(4):427–434.
- Han TU, et al. Small molecule chaperones for the treatment of Gaucher disease and GBA1-associated Parkinson disease. Front Cell Dev Biol. 2020;8:271.
-
Version history
- Version 1 (April 1, 2026): Electronic publication
Article tools
-
View PDF

- Download citation information
- Send a comment
- Terms of use
- Standard abbreviations
- Need help? Email the journal
Review Series
Neurodegeneration
-
Women’s midlife: the front line of Alzheimer prevention
Lisa Mosconi -
Genetic analysis of neurodegenerative diseases
Maurizio Grassano et al. -
Decoding neurodegeneration one cell at a time
Olivia Gautier et al. -
Lysosomal homeostasis at the crossroads of neurodegeneration
Stefano De Tito et al. -
Splicing the narrative: alternative TARDBP splicing and its relation to neurodegeneration in ALS and FTD
Morgan R. Miller et al. -
Immune signaling and function in neurodegeneration
Yvonne L. Latour et al.
Metrics
Go to
- Top
- Abstract
- Introduction
- Mechanisms of lysosomal damage and repair
- Autophagy and lysophagy
- Lysosomal dysfunction in neurodegenerative diseases
- Organelle crosstalk, system integration, and future perspectives
- Aging and network decline
- Therapeutic perspectives
- Conclusion
- Funding support
- Footnotes
- References
- Version history



Copyright © 2026 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)





