Advertisement
Commentary
Open Access |  10.1172/JCI203658
10.1172/JCI203658
The bug, the burden, and the biology: beyond host-centric phenotyping in sepsis
Georgios D. Kitsios1,2 and Rebecca M. Baron3
1Division of Pulmonary, Allergy, Critical Care and Sleep Medicine and
2Acute Lung Injury and Infection Center, University of Pittsburgh, Pittsburgh, Pennsylvania, USA.
3Division of Pulmonary and Critical Care Medicine, Department of Medicine, Brigham and Women’s Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Georgios D. Kitsios, Assistant Professor of Medicine, Division of Pulmonary, Allergy, Critical Care, and Sleep Medicine, University of Pittsburgh, Address: UPMC Montefiore Hospital, NW628, 3459 Fifth Avenue, Pittsburgh, Pennsylvania 15213, USA. Email: kitsiosg@upmc.edu.
Find articles by Kitsios, G. in: PubMed | Google Scholar
1Division of Pulmonary, Allergy, Critical Care and Sleep Medicine and
2Acute Lung Injury and Infection Center, University of Pittsburgh, Pittsburgh, Pennsylvania, USA.
3Division of Pulmonary and Critical Care Medicine, Department of Medicine, Brigham and Women’s Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Georgios D. Kitsios, Assistant Professor of Medicine, Division of Pulmonary, Allergy, Critical Care, and Sleep Medicine, University of Pittsburgh, Address: UPMC Montefiore Hospital, NW628, 3459 Fifth Avenue, Pittsburgh, Pennsylvania 15213, USA. Email: kitsiosg@upmc.edu.
Find articles by Baron, R. in: PubMed | Google Scholar
Published March 16, 2026 - More info
J Clin Invest. 2026;136(6):e203658. https://doi.org/10.1172/JCI203658.
© 2026 Kitsios, et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Related article:
Abstract
BACKGROUND Sepsis encompasses considerable biological and clinical heterogeneity. Previously, 2 phenotypes (“hyperinflammatory” and “hypoinflammatory”) have been consistently identified within sepsis via latent class analysis. These phenotypes differ in their biological features, clinical outcomes, and therapeutic responses to interventions. Prior studies of sepsis heterogeneity have focused primarily on the host response. Here, we investigate the potential influence of the causative pathogen on sepsis heterogeneity and pathobiology.METHODS We performed a retrospective observational analysis of 8,280 critically ill patients with sepsis to identify associations between pathogen characteristics and the hyperinflammatory and hypoinflammatory patient phenotypes. We also performed controlled murine and swine modeling of sepsis and lung injury and a secondary analysis of 449 patients enrolled in the EUPHRATES randomized controlled trial.RESULTS Pathogen characteristics (pathogen identity, burden, virulence, and anatomic site of infection) were strongly and independently associated with the previously reported phenotypes. In a cohort of critically ill patients with sepsis, infection with gram-negative pathogens, primarily Enterobacterales spp. (e.g., Escherichia coli, Klebsiella pneumoniae), was strongly associated with the hyperinflammatory phenotype. The hyperinflammatory phenotype was also independently associated with increased pathogen burden, virulence, and initial anatomic site of infection. In controlled murine and swine modeling, both the identity and burden of the pathogen provoked key biological features of the hyperinflammatory phenotype. Among patients with sepsis, the prognostic value of lactate clearance varied substantially by phenotype. In a secondary analysis of a randomized trial of polymyxin B hemoadsorption (which removes circulating endotoxin), hypoinflammatory patients experienced worse survival.CONCLUSIONS Our results demonstrate the central importance of pathogen features in the clinical and biological heterogeneity of sepsis. Future studies of sepsis pathobiology and heterogeneity should expand their scope beyond the host response, as understanding pathogen-host interactions will be crucial in the development of precision therapeutic strategies to improve patient outcomes.TRIAL REGISTRATION EUPHRATES trial NCT01046669.FUNDING 5P30AG024824, IK2CX002766, R01HL144599, K24HL159247, R01HL158626, R01HL173531, R35GM142992, R35GM145330, R35GM136312, K23HL166880, R35HL140026.
Authors
Rishi Chanderraj, Brian Bartek, Kathleen A. Stringer, Mohamad H. Tiba, Michael W. Sjoding, Ying He, Mark Nuppnau, Kale S. Bongers, Mark D. Adame, Sunny S. Lou, V. Eric Kerschberger, Matthew M. Churpek, Carolyn S. Calfee, Sandhya Tripathi, Debra M. Foster, John A. Kellum, Robert P. Dickson, Pratik Sinha
-
Abstract
For over a decade, sepsis phenotyping has identified hyperinflammatory and hypoinflammatory subphenotypes using host biomarkers and clinical variables, without factoring in contributions from infectious insults across patients. In this issue, Chanderraj and colleagues challenge this host-centric paradigm by demonstrating that pathogen characteristics independently contribute to sepsis subphenotypes. They reported that Enterobacterales infections, particularly Escherichia coli, strongly associated with hyperinflammatory subphenotypes, independent of illness severity. Bacterial burden, anatomic barrier breach, and circulating pathogen-associated molecular patterns influence phenotypic classification, with implications extending to culture-negative sepsis. Animal models supported causality, while reanalysis of an observational cohort and a clinical trial revealed that lactate clearance’s prognostic value and therapeutic effects of endotoxin removal with polymyxin B hemoadsorption vary by subphenotype and pathogen. These findings lay groundwork for integrative host-pathogen phenotyping; for precision medicine in critical illness, we must know not only who is sick, but what made them sick, and how the two interact.
-
Introduction
Research on critical illness and sepsis phenotyping has parsed observable heterogeneity using unsupervised methods applied to diverse data types, including transcriptomics, plasma biomarkers, and clinical variables (1–3). Despite methodological differences, recurring biological patterns emerge: hyperinflammatory states with high innate immune activation and coagulopathy; immunoadaptive versus immunosuppressed profiles; and coagulopathic/vasoplegic endotypes with endothelial injury. High-risk groups cluster around overwhelming inflammation-coagulopathy and/or impaired adaptive immunity (1). Yet these prognostically robust schemes have operated on an implicit assumption: that infectious insults are essentially equivalent across patients, with phenotypic heterogeneity arising from interindividual differences in host response to those insults (Figure 1A). The focus has been on how patients differ in response to infection, rather than on what differs in the infections themselves. What if this assumption is incomplete? What if subphenotypes also reflect different pathogens, varying in identity, burden, and virulence?
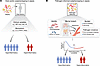 Figure 1
Figure 1Integrative host-pathogen phenotyping in sepsis: from host-centric to pathogen-informed models. (A) Traditional host-centric subphenotyping in sepsis assumes a relatively uniform microbial insult, with phenotypic heterogeneity attributed primarily to host factors. Patients are dichotomized into hyperinflammatory (red) or hypoinflammatory (blue) states based on clinical variables (such as systolic blood pressure [SBP], body temperature [temp], laboratory testing results, and heart rate [HR]) and circulating biomarkers (such as IL-6, soluble [s]TNFR1 and ANG2). Hyperinflammatory patients have higher severity of illness as measured by organ dysfunction, and worse clinical outcomes compared with hypoinflammatory patients, with potentially differential responses to treatments and interventions. (B) Pathogen-informed subphenotyping, as reported by Chanderraj et al. (4), incorporates microbial features that shape host inflammatory responses. Illustrated dimensions include pathogen identity (e.g., gram-negative rods (GNRs) such as Enterobacterales versus gram-positive cocci (GPCs) such as S. aureus); burden (e.g., time-to-positivity, lipopolysaccharide levels); barrier breach and anatomic source (i.e., bacteremia from gastrointestinal or respiratory sources); and pathogen virulence or fitness traits (e.g., fluoroquinolone-resistance [FQ-R] or -sensitivity [FQ-S]). These factors jointly influence the probability of hyperinflammatory host response, represented as a continuous spectrum rather than a binary state, with red upward arrows and blue downward arrows illustrating the impact of pathogen features on hyperinflammatory probability. In this framework, distinct pathogen characteristics shift patients toward higher or lower hyperinflammatory probabilities, emphasizing sepsis as a dynamic host-pathogen interaction rather than a fixed host-defined phenotype.
In this issue of the Journal, Chanderraj and colleagues (4) challenged this host-centric paradigm by systematically examining what has been incompletely integrated into subphenotyping frameworks: the bug, the burden, and the biology of the pathogen itself. Their findings represent a conceptual shift from viewing sepsis heterogeneity as purely host-driven toward understanding it as an integrated product of host-pathogen interactions.
-
The bug: pathogen identity matters
Chanderraj et al. analyzed 8,280 patients who were critically ill with sepsis, classifying them into subphenotypes using a validated clinical variable–based machine learning model: 22% fell into a hyperinflammatory phenotype characterized by higher illness severity (per APACHE IV scores), greater duration of shock, and higher 90-day mortality, while 78% classified as a hypoinflammatory phenotype (4). Among 1,350 patients with monomicrobial bacteremia, pathogen identity was strongly associated with subphenotype, and this association was two orders of magnitude stronger than with illness severity. Pathogen identity predicted which subphenotype a patient expressed far more consistently than prediction based on illness severity, suggesting that hyper- versus hypoinflammatory subphenotypes more strongly reflected pathogen-driven biology than host factors alone.
The specificity of pathogen identity for subphenotype was striking. Infection with Enterobacterales species — particularly E. coli and Klebsiella spp — was independently associated with the hyperinflammatory subphenotype. Patients with E. coli bacteremia had a median hyperinflammatory probability of 0.50 versus 0.14 for Staphylococcus aureus, with gram-negative bacteremia overall showing higher probabilities (0.43) than gram-positive bacteremia (0.12).
These Enterobacterales associations extended beyond bacteremia to urinary and respiratory tract infections, though with smaller effect sizes (median probabilities 0.10-0.12 vs. 0.50 for bacteremia), suggesting that barrier breach and bloodstream invasion amplify hyperinflammatory responses beyond pathogen identity alone.
-
The burden and the barrier breach
The authors demonstrated that bacterial burden (measured by time to blood culture positivity) was independently associated with hyperinflammatory classification. Shorter time to positivity, reflecting higher bacterial load, strongly predicted the hyperinflammatory subphenotype (mean 12.9 hours versus 23.9 hours, P < 0.001), with each hour of delay reducing the odds of hyperinflammatory classification (4).
Pathogen burden in blood may reflect not only the quantity of replicating organisms but also the anatomic barrier breached. Among patients with E. coli bacteremia, intraabdominal sources showed the highest hyperinflammatory probabilities, while for S. aureus bacteremia, pneumonia emerged as the highest-risk source — recapitulating observations that double-positive (blood and respiratory cultures) pneumonias portend worse outcomes (5). These patterns suggest that breach of certain anatomic barriers — particularly the gut epithelium and alveolar-capillary membrane — may permit high-burden bacteremia, amplifying the hyperinflammatory response (6).This burden-barrier framework extends beyond culture-positive cases that were the main focus of Chanderraj et al.’s study. In the EUPHRATES trial population, circulating endotoxin (lipopolysaccharide, LPS) activity correlated positively with hyperinflammatory probability, even when 70% of patients had negative blood cultures (7). Our group and others have observed that higher concentrations of plasma microbial cell–free DNA correlate with systemic inflammation, severity of organ dysfunction, and adverse outcomes in sepsis and pneumonia (8–10). Similarly, circulating β-D-glucan — a fungal cell wall component — has been associated with the hyperinflammatory subphenotype, even in patients without documented fungal infection (11). These culture-independent measures demonstrate that the amount of circulating pathogen–associated molecular patterns (PAMPs, including endotoxin, microbial DNA, β-D-glucan) can influence systemic host responses. Importantly, in culture-negative or nonbacteremic sepsis, circulating PAMPs may originate not only from a primary pathogen but also from barrier disruption that permits translocation of colonizing organisms or their fragments, emphasizing the need to study pathogen biology and mechanisms of host barrier disruption in parallel (12)
The gut warrants particular attention as a source of both live pathogens and PAMPs (13). Circulating DNA from Enterobacterales has been associated with the hyperinflammatory subphenotype (10), and intestinal domination by Enterobacterales species precedes bacteremia in immunocompromised hosts (14, 15). Host factors, including surfactant protein D expressed in the gallbladder and secreted into the gut, modulate E. coli colonization and mortality during abdominal sepsis (16), while host-pathogen interactions at the gut-luminal surface influence intestinal barrier integrity (17). This convergence of gut domination by Enterobacterales leading to barrier breach with high-burden bacteremia likely explains why intraabdominal E. coli infections associate with the highest hyperinflammatory probabilities.
-
The biology: mechanisms and clinical implications
Beyond pathogen identity and burden, Chanderraj et al. examined virulence as a determinant of subphenotypes (4). Using fluoroquinolone resistance as a proxy for fitness cost in Enterobacterales species, they found that fluoroquinolone-resistant isolates — which exhibit reduced virulence in vitro — were associated with decreased probability of the hyperinflammatory subphenotype (0.14 versus 0.57, P < 0.001). This observation underscores that not all Enterobacterales infections are immunologically equivalent. More broadly, the biological basis for Enterobacterales’ association with hyperinflammation may involve LPS, a potent TLR4 activator whose structure — particularly the lipid A moiety — can modulate immunogenicity by up to several orders of magnitude (18).
On the host side, hyperinflammatory probability correlated more strongly with procalcitonin, aligning with prior latent class analyses (19). Importantly, lactate clearance (20) conferred stronger survival benefit in hyperinflammatory patients, with similar effects in Enterobacterales bacteremia. Chanderraj et al.’s reanalysis of the EUPHRATES trial revealed significant treatment effect modification; specifically, Polymyxin B hemoadsorption — which removes endotoxin — was associated with statistically significant harm in patients who were hypoinflammatory, while showing no benefit in patients who were hyperinflammatory (4). Together, these findings suggest that pathogen features impact inflammatory intensity and host metabolism; however, predicting treatment response may ultimately require integration of both pathogen characteristics and host response profiling.
Controlled animal models corroborated these observations (4). In swine, E. coli sepsis with direct lung injury provoked higher hyperinflammatory probabilities compared with lung injury alone. In murine peritonitis using two Enterobacterales species, increasing bacterial inoculum dose-dependently elevated inflammatory biomarkers (sTNFR1, IL-6, CXCL1). While these models have inherent limitations, including use of artificial inocula, direct organ inoculation, and species differences, they support the directionality of the authors’ thesis: pathogen characteristics causally influence the development of hyperinflammatory responses. These and other translational studies remain essential to understanding host-pathogen heterogeneity and identifying therapeutic targets (16, 21), even if they cannot fully recapitulate the complexity of human sepsis.
Despite the meaningful findings presented in this study, mechanism remains incompletely defined, and questions remain. Do differences in LPS structure explain the specificity observed? Does host genetic variation in pattern recognition receptors modulate responsiveness? Are pathogen characteristics altered by interaction with host-derived proteins? Are other PAMPs, such as flagellin or peptidoglycan, contributing? What is the influence of the microbiome on specific pathogens and host responses upon presentation and following broad-spectrum antibiotic therapy or stress-dose steroids? Future studies integrating microbiota profiles, dominant pathogen genomics, host receptor genetics, and functional immunology will be essential. Notably, pathogen-informed subphenotypes may represent mechanistically distinct endotypes, i.e. biological disease variants defined by pathogen-specific triggering of host pathways. If E. coli–driven hyperinflammation operates through different molecular mechanisms than S. aureus–driven responses, these represent true endotypes rather than simply prognostic clusters, with implications for targeted therapeutic development.
-
The boundaries: limitations of pathogen-informed subphenotyping in sepsis
This work has important limitations that define future priorities. First, culture-positive bacteremic sepsis represents only 10%–15% of community-onset sepsis cases (22). While endotoxin analysis and prior culture-independent microbial profiling work suggest these findings generalize to culture-negative sepsis (8–10), prospective validation integrating both culture-based and culture-independent diagnostics is needed (9, 21).
Second, pathogen characteristics explain only modest variance in subphenotype classification. Host factors such as genetics, comorbidities, immune history, and microbiome composition may also be important. Pathogen features are not deterministic; rather, they act as proximal triggers whose effects depend on host capacity to contain or amplify inflammation. Moreover, pathogens interact with complex microbial communities, such that sepsis often reflects polymicrobial dynamics rather than single-pathogen biology. (12).
Third, clinical variable–based classifiers may have up to 22% misclassification versus biomarker models (23). While clinical models offer accessibility, targeted immunomodulation trials may require the biological fidelity of biomarker models.
Fourth, these subphenotypes evolve dynamically, with hyperinflammatory probabilities declining over the ICU course (24, 25). Whether these transitions reflect pathogen clearance, or whether persistent or recurrent infections prevent recovery, remains unknown. Longitudinal studies integrating microbial burden with subphenotype trajectories could identify optimal timing for interventions.
Finally, mechanistic pathways require further interrogation. Beyond PAMPs and pattern recognition receptors, pathogen-derived metabolites may shape host responses: acetylated polyamines produced by gram-negative bacteria during bloodstream infection influence both bacterial virulence and host metabolism (21). Addressing these questions requires deeply phenotyped cohort studies with serial host and pathogen sampling, particularly from abdominal sources of sepsis. The majority of accumulated cohorts reflect predominantly respiratory sepsis (26), yet, abdominal-derived pathogens appear critical to hyperinflammatory responses (4).
-
The bold road ahead: integrative host-pathogen phenotyping
Chanderraj and colleagues establish that pathogen characteristics — identity, burden and virulence — are independent predictors of sepsis subphenotypes, moving beyond the assumption that subphenotypes reflect only host heterogeneity (Figure 1B). Future clinical trials should integrate pathogen diagnostics and host responses into enrollment and stratification schemes. Platform trials, such as the PANTHER network for acute respiratory distress syndrome (ARDS) (27), offer a framework for testing subphenotype-targeted and pathogen-informed interventions simultaneously. To realize precision medicine in critical illness, we need to know not only who is sick, but what made them sick, and how the two interact. This study brings us meaningfully closer to that goal.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- NIH (R01 176668; to GDK).
-
Footnotes
Conflict of interest: GDK has received research funding from Pfizer Inc., and Genentech Inc., unrelated to this work, consulting fees from InflaRx Inc., unrelated to this work, and serves on the Advisory Board of KeepBio.
Copyright: © 2026, Kitsios et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: J Clin Invest. 2026;136(6):e203658. https://doi.org/10.1172/JCI203658.
See the related article at Pathogen characteristics are key determinants of distinct host response phenotypes of sepsis.
-
References
- Shankar-Hari M, et al. Reframing sepsis immunobiology for translation: towards informative subtyping and targeted immunomodulatory therapies. Lancet Respir Med. 2024;12(4):323–336.
- Sinha P, et al. Identifying molecular phenotypes in sepsis: an analysis of two prospective observational cohorts and secondary analysis of two randomised controlled trials. Lancet Respir Med. 2023;11(11):965–974.
- Seymour CW, et al. Derivation, validation, and potential treatment implications of novel clinical phenotypes for sepsis. JAMA. 2019;321(20):2003–2017.
- Chanderraj R. Pathogen characteristics are key determinants of distinct host response phenotypes of sepsis. J Clin Invest. 2026;136:e197346.
- Haessler S, et al. Blood cultures versus respiratory cultures: 2 different views of pneumonia. Clin Infect Dis. 2020;71(7):1604–1612.
- Haidar G, et al. Emergence of high-level aztreonam-avibactam and cefiderocol resistance following treatment of an NDM-producing Escherichia coli bloodstream isolate exhibiting reduced susceptibility to both agents at baseline. JAC Antimicrob Resist. 2024;6(5):dlae141.
- Dellinger RP, et al. Effect of targeted polymyxin B hemoperfusion on 28-day mortality in patients with septic shock and elevated endotoxin level: the EUPHRATES randomized clinical trial. JAMA. 2018;320(14):1455–1463.
- Yang H, et al. Circulating microbial cell-free DNA is associated with inflammatory host-responses in severe pneumonia. Thorax. 2021;76(12):1231–1235.
- Lisius G, et al. Noninvasive diagnosis of secondary infections in COVID-19 by sequencing of plasma microbial cell-free DNA. iScience. 2023;26(11):108093.
- Neyton LPA, et al. Host and microbe blood metagenomics reveals key pathways characterizing critical illness phenotypes. Am J Respir Crit Care Med. 2024;209(7):805–815.
- Kitsios GD, et al. Plasma 1,3-β-d-glucan levels predict adverse clinical outcomes in critical illness. JCI Insight. 2021;6(14):e141277.
- Kitsios GD, Bain W. “Now we got bad blood”: beyond phenotype labels in an “era” of meta-omics in critical illness. Am J Respir Crit Care Med. 2024;209(7):772–774.
- Nath S, et al. Gut-lung crosstalk during critical illness. Curr Opin Crit Care. 2023;29(2):130–137.View this article via: PubMed Google Scholar
- Shimasaki T, et al. Increased relative abundance of Klebsiella pneumoniae carbapenemase-producing Klebsiella pneumoniae within the gut microbiota is associated with risk of bloodstream infection in long-term acute care hospital patients. Clin Infect Dis. 2019;68(12):2053–2059.
- Taur Y, et al. Intestinal domination and the risk of bacteremia in patients undergoing allogeneic hematopoietic stem cell transplantation. Clin Infect Dis. 2012;55(7):905–914.
- Varon J, et al. Surfactant protein D influences mortality during abdominal sepsis by facilitating Escherichia coli colonization in the gut. Crit Care Explor. 2022;4(5):e0699. View this article via: PubMed Google Scholar
- Matute JD, et al. Intelectin-1 binds and alters the localization of the mucus barrier-modifying bacterium Akkermansia muciniphila. J Exp Med. 2023;220(1):e20211938.
- Needham BD, et al. Modulating the innate immune response by combinatorial engineering of endotoxin. Proc Natl Acad Sci U S A. 2013;110(4):1464–1469.
- Drohan CM, et al. Biomarker-based classification of patients with acute respiratory failure into inflammatory subphenotypes: a single-center exploratory study. Crit Care Explor. 2021;3(8):e0518. View this article via: PubMed Google Scholar
- Hernández G, et al. Effect of a resuscitation strategy targeting peripheral perfusion status vs serum lactate levels on 28-day mortality among patients with septic shock: the Andromeda-shock randomized clinical trial. JAMA. 2019;321(7):654–664.
- Mayers JR, et al. A metabolomics pipeline highlights microbial metabolism in bloodstream infections. Cell. 2024;187(15):4095–4112.
- Ohnuma T, et al. Epidemiology, resistance profiles, and outcomes of bloodstream infections in community-onset sepsis in the United States. Crit Care Med. 2023;51(9):1148–1158.
- van Amstel RBE, et al. Open-source clinical classifiers for ARDS: democratizing precision or diluting biology? Intensive Care Med. 2025;51(11):2123–2125.
- Van Amstel RBE, et al. Temporal transitions of the hyperinflammatory and hypoinflammatory phenotypes in critical illness. Am J Respir Crit Care Med. 2025;211(3):347–356.
- Lu M, et al. Trajectories of host-response subphenotypes in patients with COVID-19 across the spectrum of respiratory support. CHEST Crit Care. 2023;1(3):100018.
- ARDS. ARDS, Pneumonia, and Sepsis (APS) Consortium. https://apsconsortium.org/ Accessed February 4, 2026.
- Reddy K, et al, eds. Building an international precision medicine platform trial for the acute respiratory distress syndrome (ARDS): an expert consensus project report. National Institute for Health and Care Research; 2025.
-
Version history
- Version 1 (March 16, 2026): Electronic publication
Article tools
-
View PDF

- Download citation information
- Send a comment
- Terms of use
- Standard abbreviations
- Need help? Email the journal
Metrics
Go to
- Top
- Abstract
- Introduction
- The bug: pathogen identity matters
- The burden and the barrier breach
- The biology: mechanisms and clinical implications
- The boundaries: limitations of pathogen-informed subphenotyping in sepsis
- The bold road ahead: integrative host-pathogen phenotyping
- Funding support
- Footnotes
- References
- Version history



Copyright © 2026 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)

