Advertisement
Review
Open Access |  10.1172/JCI205086
10.1172/JCI205086
Towards precision medicine for brain arteriovenous malformations
Andrew T. Hale,1,2 Adam J. Kundishora,3 Pazhanichamy Kalailingam,4 Tanyeri Barak,5 Phan Q. Duy,6 Christopher M. Ramundo,6 Baojian Fan,6 Qiang Li,6 Priscilla K. Brastianos,7 Ganesh M. Shankar,6 Seth L. Alper,7,8,9 Benjamin P. Kleinstiver,4,10,11 Patricia L. Musolino,4,12 and Kristopher T. Kahle6,9,13,14
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by
Hale, A.
in:
PubMed
|
Google Scholar
|

1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Kundishora, A. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Kalailingam, P. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Barak, T. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Duy, P. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Ramundo, C. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Fan, B. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Li, Q. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Brastianos, P. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Shankar, G. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Alper, S. in: PubMed | Google Scholar
1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by
Kleinstiver, B.
in:
PubMed
|
Google Scholar
|

1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by
Musolino, P.
in:
PubMed
|
Google Scholar
|

1Department of Neurosurgery, University of Alabama at Birmingham, Birmingham, Alabama, USA.
2Neuroscience Institute, University of Cape Town, Cape Town, South Africa.
3Division of Neurosurgery, Children’s Hospital of Philadelphia, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania, USA.
4Center for Genomic Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
5Department of Neurosurgery, Yale University School of Medicine, New Haven, Connecticut, USA.
6Department of Neurosurgery, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
7Department of Medicine, Massachusetts General Hospital, Boston, Massachusetts, USA.
8Division of Nephrology and Vascular Biology Research Center, Beth Israel Deaconess Medical Center, Boston, Massachusetts, USA.
9Broad Institute of MIT and Harvard, Cambridge, Massachusetts, USA.
10Department of Pathology, Massachusetts General Hospital, Boston, Massachusetts, USA.
11Department of Pathology, Harvard Medical School, Boston, Massachusetts, USA.
12Department of Neurology, Harvard Medical School and Massachusetts General Hospital, Boston, Massachusetts, USA.
13Division of Genetics and Genomics, Boston Children’s Hospital, Boston, Massachusetts, USA.
14Harvard PhD Program in Neuroscience (PiN), Harvard University, Cambridge, Massachusetts, USA.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
Find articles by Kahle, K. in: PubMed | Google Scholar
Published May 15, 2026 - More info
J Clin Invest. 2026;136(10):e205086. https://doi.org/10.1172/JCI205086.
© 2026 Hale et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
-
Abstract
Recent advances in cerebrovascular genomics, single-cell biology, pharmacology, and gene editing technology are transforming our understanding of brain arteriovenous malformations (bAVMs) — a leading cause of pediatric hemorrhagic stroke. Once considered static anatomical defects, bAVMs are now recognized as dynamic, genetically driven lesions resulting from somatic mutations in KRAS, BRAF, and pathways involved in arteriovenous specification, angiogenesis, and vascular remodeling. By integrating human genetics, animal models, and endovascular innovations, researchers have uncovered convergent mechanisms that link endothelial Ras/MAPK hyperactivation to abnormal vessel growth and higher rupture risk. These insights provide a foundation for precision medicine approaches that combine molecular diagnostics — such as liquid or endoluminal biopsies — with mutation-specific pharmacotherapies and emerging CRISPR-based gene editing strategies. We suggest that genotype-guided interventions, tailored by spatial and developmental cerebrovascular context, could ultimately reclassify bAVMs from surgically incurable malformations to treatable molecular conditions.
-
Introduction
Brain arteriovenous malformations (bAVMs) are direct connections between arteries and veins without an intervening capillary bed, creating high-flow, low-resistance vascular circuits (1). Estimates for sporadic bAVM incidence are approximately 0.9–1.4 per 100,000 person-years, with a prevalence of about 5–18 per 100,000 (2, 3). bAVM rupture is a leading cause of atraumatic hemorrhagic stroke in children and can result in severe morbidity, permanent neurologic disability, or death. Treatment options include microneurosurgical resection or stereotactic radiosurgery (SRS), which may involve adjunctive endovascular embolization treatments. However, many bAVMs remain untreatable due to their large size, eloquent location, and/or lack of response to SRS, among other factors. The frequent diagnosis of bAVMs in childhood and an annual rupture rate of approximately 4% (4) highlight the need for new treatment paradigms to attenuate risk of bAVM rupture. Repurposed molecularly targeted pharmacotherapeutics and/or gene editing therapies (i.e., CRISPR/Cas nucleases, prime editors, or base editors; refs. 5–7) could benefit the approximately 50%–80% of patients with sporadic KRAS/BRAF-mutant bAVMs (8–10) as well as patients with a bAVM-associated syndrome, where an inherited or (rarely) de novo germline mutation in the presence of a somatic “second hit” causing biallelic loss or environmental trigger (angiogenic stimuli, shear stress, transzygosity, etc.) defines AVM pathogenesis.
Here we detail the molecular mechanisms, human genetics, animal models, translational studies, and key clinical research supporting the rationale for pharmacologic and/or gene editing therapies for bAVMs. We propose a mechanistic framework for bAVM precision therapies based on our understanding of (a) signaling pathways that control angiogenesis, arteriovenous (AV) specification and zonation, and cerebrovascular (CV) cell identity; (b) spatiotemporal variations in CV single-cell developmental pathways; (c) abnormal endothelial growth, response to growth factors, and radiosensitivity; (d) animal models and human genetics related to sporadic and syndromic bAVM development; (e) common pathogenic mutations linking sporadic bAVMs and cancers with mutant-specific and pathway-modulatory therapies; and (f) tailored gene editing technologies and delivery methods that offer potential for a single, permanent correction. These insights lay the groundwork for genotype-directed precision medicine for select patients with bAVMs.
-
Clinical overview of bAVMs
bAVMs present across a clinically heterogeneous spectrum with variable risk of hemorrhage, epilepsy, neurologic deficit, and systemic involvement that arise in distinct phenotypic and genetic contexts. Hereditary hemorrhagic telangiectasia (HHT1, -2, and -5) is caused by autosomal dominant inherited, or rarely de novo, mutations in ALK1/ACVRL1, ENG, GDF2, or SMAD4 (11, 12), where somatic biallelic loss or an environmental trigger (i.e., angiogenic stimulus) defines AVM pathogenesis. HHT is typically diagnosed based on family history and identification of systemic manifestations (low-flow mucocutaneous telangiectasias, capillary malformations, and/or epistaxis) that appear in adolescence or adulthood, with bAVMs occurring in approximately 20% and 5% of HHT1 and HHT2 patients, respectively. In contrast, patients with CM-AVM — caused by somatic second-hit loss-of-function mutations in the setting of heterozygous germline mutations in RASA1 and EPHB4 (13) — typically present at birth or early infancy. Patients with CM-AVM have diffuse, halo-ring capillary malformations in the trunk/limbs and angioarchitecturally distinct, high-flow brain vascular malformations, including vein of Galen malformations (VOGMs), AV fistulas, or less commonly true parenchymal bAVMs. Sporadic VOGMs can present in the neonatal, infant, or early childhood periods with high-output cardiac failure, macrocephaly, and/or hydrocephalus and is associated with high morbidity and mortality. Approximately 20% of cases are caused by de novo or inherited germline variants in EPHB4, RASA1, ACVRL1, NOTCH1, ITGB1, and PTPN11, and postzygotic second-hit somatic mutations are hypothesized to define the remaining cases (14). Sturge-Weber syndrome is caused by somatic mutations in GNAQ and presents with facial capillary malformations (i.e., port wine stain) and seizures, secondary to hemispheric leptomeningeal vascular malformations (without formation of a true bAVM nidus), causing progressive neurologic impairment (15). In contrast, sporadic bAVMs are solitary lesions caused by somatic activating mutations in KRAS, and less commonly BRAF, that can present across the age spectrum with hemorrhage, seizures, headache, focal neurologic deficits, or incidentally where clinical manifestations are determined by variable angioarchitectural features, including size, anatomical location, and pattern of venous drainage, among others. We hypothesize that integrating knowledge of genetic drivers, molecular mechanisms, and clinical behavior in the context of angioarchitectural features across this bAVM spectrum will enable tailoring precision therapeutic strategies.
-
Angiogenesis, AV specification, zonation, and CV development
CV growth, development, and remodeling occur through angiogenesis, either by sprouting from existing vessels or by partitioning into daughter branches via intussusception. These processes depend on tightly regulated molecular, genetic, and environmental cues (16). Both angiogenesis mechanisms involve proliferation, migration, patterning, cellular renewal, and morphological remodeling. Genetic and environmental modulators of angiogenesis — including shear stress (17), inflammation (18) and radiation (19) — shape CV identity, function, connections, and mechanical stability. Although multiple angiogenic phenotypes result from genetic alteration or pharmacologic manipulation of these pathways, we anticipate that specific knowledge of signaling pathways responsible for bAVM growth, maintenance, and regression will enable molecularly guided precision treatments. Aberrant spatial or temporal expression of angiogenic factors underlies bAVM histopathology (20), and risk variants in angiogenesis-associated genes may influence bAVM formation and hemorrhage (21). Therefore, framing molecularly guided bAVM treatment requires an overview of pathways regulating angiogenesis, AV specification, zonation, and CV development.
Ephrin/Eph/RASA1. The ten EphA receptor isoforms and six EphB receptor isoforms are receptor tyrosine kinases (RTKs) that coordinate bidirectional signaling through direct interactions with the five ephrin A ligand isoforms and three ephrin B isoforms to regulate angiogenesis (Figure 1) and other cellular processes (22). Exquisite control, crosstalk, and pleiotropy among members of the ephrin/Eph pathway control endothelial cell (EC) migration and adhesion and AV differentiation, specification, and zonation in normal and pathological CV (23). Expression of ephrin B2 and EphB4 define arterial and venous ECs, respectively, during development (24). This pattern is maintained (25) but the ratio of cell types is perturbed in human bAVMs (26). Whether this perturbation is a consequence of bAVM formation or a primary mechanistic driver of bAVM pathogenesis, growth, and/or maintenance is not known. Germline SNPs in EPHB4 have been associated with rupture risk in sporadic adult bAVMs (27), but the mechanistic basis for this observation is not clear. Our group and others have identified causative EPHB4 loss-of-function mutations in a subset of human patients with the congenital/pediatric bAVM subtype VOGM (14, 28, 29). Heterozygous mice expressing a VOGM-specific Ephb4 kinase domain missense variant (Ephb4F867L) displayed severe CV abnormalities only in the presence of a second variant allele or a Cre-disrupted Ephb4 allele (14).
 Figure 1
Figure 1Overview of ephrin/Eph/RASA1, PI3K/AKT/mTOR, VEGF, NOTCH, TGF-β/SMAD, BRAF, and KRAS pathways. These signaling pathways underlie angiogenesis, AV specification, and CV development. Eph, ephrin; ACVRL1/ALK1, activin receptor–like kinase 1; SMAD, suppressor of mothers against decapentaplegic; SARA, SMAD anchor for receptor activation; Smurf2, SMAD ubiquitin regulatory factor 2; VRAP, VEGF receptor–associated protein; Shc, Src-like kinase C; PTEN, phosphatase and tensin homolog deleted on chromosome 10; AKT, protein kinase B; mTORC1, mammalian target of rapamycin complex 1; PTPN1, tyrosine-protein phosphatase non-receptor type I; RASA1, p120 Ras GTPase–activating protein; RAF, rapidly accelerated fibrosarcoma.
Downstream activation of RAS suppressor p120 Ras GTPase–activating protein (RASA1), a negative regulator of RAS signaling, is the mechanism through which EphB4 mutations regulate angiogenesis. EphB4 and RASA1 physically interact via an SH2 domain in RASA1 and a phosphorylated tyrosine residue in EphB4 (30) to limit RAS signaling in ECs (14). RAS activation also causes downstream activation of VEGF-dependent signaling (31) and is sufficient to cause bAVMs in mice (32). Somatic RASA1 variants cause capillary malformation AVM syndrome (CM-AVM) (33, 34), characterized by high-flow vascular malformations, including systemic and intracranial AV fistulas, and, rarely, bAVM subtype VOGM (35, 36). Moreover, mice with an Ephb4 mutation that prevents physical interaction with RASA1 but retains protein tyrosine kinase activity show normal angiogenic phenotypes (37). These data suggest, but do not prove, that EphB4 and RASA1 act within the same pathway to regulate angiogenesis. Pharmacological manipulation of RASA1 or downstream activators (e.g., mTOR) may be an appealing approach to treat genetically susceptible bAVM subtypes. However, somatic activating mutations in EPHB4 and RASA1 have not been identified in human patients with sporadic bAVMs.
PI3K/AKT/mTOR. Phosphoinositide 3-kinases (PI3Ks) are lipid kinases (class I, II, and III) that control downstream signaling pathways, including the serine-threonine kinases AKT and mTOR to control cell growth, proliferation, migration, and survival in the context of angiogenesis (Figure 1). Ligand-receptor binding or environmental stimuli such as shear stress (17) promote phospholipase C–mediated cleavage of phosphoinositide 4,5-bisphosphate (PIP2) to generate the second messengers diacylglycerol (DAG) and inositol-1,4,5-triphosphate (IP3) (38). Somatic gain-of-function mutations in PI3KCA have been identified in venous malformations (39) and cerebral cavernous malformations (40) in mice and humans, but not sporadic bAVMs or other bAVM subtypes. Pharmacologic inhibition of PI3K or mTOR leads to regression of PI3K-driven vascular malformations (41) and is therapeutic in mouse models of HHT (42, 43). However, in mice harboring EC-specific HrasG12V mutations, PI3K inhibition had no effect in vivo on AVM-like features (i.e., vascular dysplasia, hemorrhage, etc.) in the brain and other organs (32). Of note, to our knowledge, somatic activating mutations in HRAS, NRAS, or PI3K/AKT/mTOR pathway genes have not been identified in human patients with sporadic bAVMs. Importantly, inhibition of MEK, but not PI3K, signaling can abrogate angiogenesis through VEGF-dependent pathways in human KRAS-mutant bAVM ECs (10). Thus, pharmacotherapies against PI3K/AKT/mTOR are unlikely to provide therapeutic benefit for patients with sporadic bAVMs.
VEGF. The VEGFA-VEGFR2 interaction is the main signaling pathway driving CV angiogenesis, although other VEGF ligands (VEGFB–D) and receptors (VEGFR1–3) also have context-specific roles (Figure 1). VEGFA signaling has been extensively studied in bAVM pathogenesis in preclinical models, where overactivation of VEGFA causes CV dysmorphogenesis (44) and hemorrhage that can be attenuated by MMP9 inhibition (45). Additional support for VEGF signaling in bAVM pathogenesis is provided by mouse studies demonstrating that VEGF inhibition via treatment with the monoclonal antibody bevacizumab attenuates vascular dysplasia in mature bAVMs of Alk1-deficient mice (46). In contrast, two patients (genotype unknown) with sporadic bAVM treated with bevacizumab did not experience reduced bAVM volume, although the reduced VEGF serum levels observed at 26 weeks of treatment were not sustained through the end of the 52-week treatment period (47). Although no adverse systemic effects (hypertension, proteinuria, impaired wound healing, etc.) (48) of bevacizumab were reported in this case series, additional safety data are needed. In one series of patients with sporadic bAVMs, germline variants were identified in genes involved in VEGF and BMP/TGF-β pathways (49), but clear pathogenic evidence for these variants is lacking. Moreover, VEGF signaling can be augmented by shear stress (50) and iatrogenic bAVM embolization (51) through disruption of the blood-brain barrier (BBB) (52). These data highlight the highly dynamic regulation of VEGF in vivo, and suggest underlying mechanisms by which VEGF contributes to bAVM biology and treatment response. We hypothesize that VEGF signaling contributes to bAVM pathogenesis via augmentation of vascular stability and remodeling rather than as a primary genetic driver. While there is some compelling preclinical evidence, limited and variable human outcomes data, along with potential systemic toxicities, tempers enthusiasm for VEGF as a therapeutic target.
TGF-β/SMAD. TGF-β/SMAD signaling has both pro- and antiangiogenic roles. Clinical studies of HHT confirmed that loss-of-function variants in endoglin (ENG) (HHT1) or ALK1/ACVRL1 (HHT2), both members of the TGF-β superfamily, cause the disease (53, 54). For example, ALK1/ACVRL1 is a receptor that, when bound by a ligand, phosphorylates SMAD1/5/8, enabling the latter to form a complex with SMAD4 (mutated in HHT5) and translocate to the nucleus (Figure 1). Loss of ALK1/ACVRL1 removes the angiogenic “brake” on VEGF-mediated PI3K signaling, leading to abnormal sprouting, EC proliferation, and vascular dysplasia (55). Loss-of-function mutations in Eng, a transmembrane coreceptor for ALK1/ACVRL1, cause bAVMs through similar mechanisms (56–58). Mouse models lacking systemic or EC-specific Smad4 exhibit systemic AVMs and bAVMs that respond to angiopoietin-2 inhibition (59–63). Furthermore, decreased expression of TGF-β/BMP and downstream SMAD6 has been linked to microhemorrhage in unruptured human sporadic bAVM surgical samples (64). Thus, it appears that changes in TGF-β/SMAD signaling, whether directly or via upstream regulation, may contribute to the development of bAVM pathogenesis through complex regulation of angiogenesis.
Notch. Notch signaling occurs through ligand-receptor interactions (DLL1/3/4, JAG1/2 binding NOTCH1–4), resulting in γ-secretase–dependent release of the Notch intracellular domain (NICD) and nuclear translocation to regulate transcription via CBF-1/RBPJ (Figure 1). Modulation of NOTCH1, -3, and -4 and RBPJ — leading to either increased or decreased Notch signaling — has been implicated in bAVM, extracranial AVM, and vascular dysgenesis in mice (65–73), but no pathogenic NOTCH mutations have been identified in human sporadic bAVMs. Interpretation of Notch-dependent phenotypes is complicated by downstream signaling redundancy, crosstalk, varied transcriptional programs, cell-type specificity, sequestration of Notch binding partners, differential ligand binding, and compensatory mechanisms (74). For example, deletion of the downstream Notch effector gene Rbpj has been shown to both cause vascular dysmorphogenesis and also rescue NOTCH4-dependent AVM formation (71–73), whereas Notch deficiency in pericytes causes extracranial AVMs and bAVMs (75). Mice lacking EphB4 or Efnb2 feature embryonic vascular defects similar to those observed in mice harboring Notch1 gain-of-function mutations (67), but these phenotypes do not resemble sporadic, adult-onset bAVMs. Germline NOTCH4 variants have also been suggested to increase risk of human bAVM formation (65), but independent replication and direct causal evidence is lacking. Additional data support shear stress– and vascular tone–mediated regulation of Notch contributing to bAVM pathogenesis (76, 77), and the therapeutic effect of radiation on bAVMs may be mediated in part by Notch signaling (78). Although Notch perturbation in mice rarely results in bAVM phenotypes independently from angiogenic modulation, these studies have been instrumental in identifying the essential role of Notch signaling in vascular development. The overlapping and convergent mechanisms with ephrin and TGF-β/SMAD suggest that these pathways may be modulated in specific bAVM biological states and in response to radiation. Thus, the role of Notch in human sporadic bAVM pathogenesis is likely dependent on genotype and environment.
KRAS. KRAS, a GTPase downstream of RTKs, is a major cellular regulator of growth and proliferation and has been extensively studied in oncology. The discovery of somatic mutations in KRAS underlying approximately 50%–80% of sporadic bAVMs (8–10) has directed attention to the role of KRAS as an angiogenesis regulator (Figure 1). KRAS activation has been shown to augment endothelial-mesenchymal transition (endoMT) through upstream modulation of TGF-β/SMAD, through a mechanism attenuated by lovastatin (79). KRAS-activating mutations cause downstream activation of PI3K-independent MAPK/ERK signaling (80). The prevalence of KRAS gain-of-function mutations in sporadic bAVMs suggests that anti-VEGF therapy may be unsuccessful, although as noted above there are very limited bevacizumab bAVM treatment data in human patients (47). KRAS-mutant-specific pharmacology, downstream MEK inhibition, and primary correction of pathogenic somatic mutations (or KRAS hyperactivation) may be viable nonsurgical treatments for human sporadic bAVMs, but additional mechanistic studies coupled with rigorous clinical trials and safety data are needed.
-
Genetic basis of bAVM pathogenesis
Human genetics of syndromic bAVMs. Syndromes associated with bAVM development — caused by autosomal dominant inherited, or less commonly, de novo variants — include HHT1, -2, and -5 (caused by mutations in ENG, ALK1/ACVRL1, and SMAD4, respectively) (53, 54), CM-AVM (RASA1 and EPHB4 mutations) (13, 81), Parkes-Weber syndrome (RASA1 mutations) (35), and Sturge-Weber syndrome (somatic GNAQ mutations) (82) (Table 1). However, most cases are incompletely penetrant and variably expressive, leading to AVMs in different locations (brain, lung, liver, etc.), where somatic second-hit loss-of-function mutations or triggers (i.e., transzygosity in modifier genes, shear stress, angiogenic stimuli, etc.; ref. 83) have been shown to determine the spatiotemporal parameters of AVM pathogenesis. Accepted mechanisms causing bAVMs in HHT include changes in PI3K activation and aberrant Notch signaling (55, 84), whereas CM-AVM cases caused by EPHB4 and RASA1 mutations lead to aberrant activation of RAS signaling (13, 81). However, these bAVMs differ pathologically. HHT-associated bAVMs are typically lower-flow and characterized by vessel dysplasia (more common in patients with HHT1 [~20%] than in those with HHT2 [5%]), with systemic clinical signs (i.e., mucocutaneous capillary malformations) typically appearing in adolescence or adulthood. In contrast, CM-AVM–related brain and other AVMs are high-flow and congenital or infantile, including VOGM, AV fistulas, or rarely parenchymal bAVMs that are pathologically distinct from sporadic bAVMs. Therefore, precision medicine approaches must account for the distinct genetic drivers, mechanisms, and pathophysiological characteristics of syndromic bAVMs.
Human genetics of sporadic bAVMs. Pathogenic somatic mutations in KRAS, and less frequently BRAF, have been found in approximately 50%–80% of surgically resected, sporadic human bAVMs (8–10). A summary of human genetic studies on sporadic bAVMs is shown in Table 2. Nikolaev et al. first identified the role of somatic activating mutations in KRAS (G12D, G12V, and Q61H) (10). These mutations are found at low allele frequencies (0.5%–4%), are localized to ECs, and lead to MAPK/ERK hyperactivation. Reproduction of somatic KRAS mutations (G12D and G12V) and the discovery of new variants (G12A, G12C, and S65_A66insDS) were reported in separate human cohorts (8, 9, 85), in which G12A and G12C were also identified in patients with spinal AVMs (9).
Germline SNPs and de novo variant (DNV) analysis of human bAVMs reveal additional possible genetic contributions, but the causative evidence for these variants is lacking. For example, germline polymorphisms in angiopoietin-like 4 (ANGPTL4) (86) and DNVs across 46 genes converging on endoMT and on TGF-β/SMAD and VEGF signaling pathways have been implicated in bAVM risk (49, 87). Moreover, DNV analysis of 152 sporadic bAVM proband-parent trios and 40 singletons implicated multiple genes, including ANGPTL3 and SLC19A3 (88), whereas an independent cohort of 60 sporadic bAVM proband-parent trios identified nonsynonymous germline variants in 46 genes enriched in vascular cell types and involved in endoMT, including EXPH5, EPAS1, and ENG (87). Finally, in a series of 112 additional sporadic bAVM proband-parent trios, compound heterozygous variants in 16 genes were implicated in more than one trio, including five genes (MYLK, HSPG2, PEAK1, PIEZO, and PRUNE2) associated with angiogenesis or vascular disease (89). However, most of these variants have not been replicated or mechanistically validated (Table 2).
-
Single-cell analysis of the cerebrovasculature and bAVMs
Construction of large-scale cellular and transcriptomic atlases highlights the diversity, complexity, and heterogeneity of normal human CV system across development and disease (90–93). However, critical limitations to single-cell and spatial approaches in human bAVMs include surgical sampling bias to nidal tissue without parenchyma, reliable detection of low-frequency mutant ECs, lack of hemodynamic context, and a cross-sectional analysis without temporal developmental insights. Nonetheless, recent single-cell RNA sequencing (scRNA-seq) and supporting molecular validation experiments have identified changes within the CV in bAVMs, including accumulation of arterially fated and venous cluster ECs, presumably due to loss of capillary EC populations (a hallmark histopathological feature of bAVMs) (92). Mechanistically, Wälchli et al. demonstrated reactivation of a fetal gene signature, including multiple angiogenic pathways and angiogenic capillary markers PVLAP (and ESM1) that may be reflective of PVLAP’s role as an immature and non–BBB-competent EC marker (94). Moreover, these authors demonstrate disruption of the BBB and brain EC identities underscoring bAVM pathogenesis (92). Additional data in KrasG12D-driven bAVMs in mice also suggests that an increase in the tip cell population and angiogenic markers may be driven by androgen- and Notch-dependent signaling, providing rationale for investigation of therapeutic modulation of these pathways (95). An independent single-cell analysis of 106,853 cells across five resected bAVM specimens revealed 11 unique cell types, an increase in angiogenic potential and endoMT within the nidus, a unique immune cell signature including recruitment of myeloid cells after bAVM rupture, and modulation of hemorrhage risk via immune-mediated crosstalk with mural cells (93). Independent scRNA-seq in 17 resected bAVMs identified three unique EC clusters and one mural/fibroblast cluster (87). Finally, differential CpG methylation in EPHB1 and KRAS (among other genes) has been observed in human bAVM ECs (96). Taken together, these single-cell data frame bAVMs as lesions driven by EC identity, augmented AV developmental trajectories, BBB disruption, and pathologic inflammation coordinating EC–mural cell crosstalk contributing to rupture pathogenesis (Figure 2). Future efforts to resolve the mechanistic consequences of mosaic patterning of mutant cells and integration of bAVM genotype-phenotype molecular programs will inform therapeutic strategies.
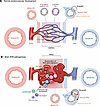 Figure 2
Figure 2Single-cell architecture of normal CV development and bAVM pathogenesis. (A) Normal CV development is characterized by carefully choreographed arterial-capillary-venous development through coordinated angiogenic sprouting and endothelial specificity. Mesenchymal and vascular smooth muscle cells (SMCs) provide structural support. (B) Pathogenic bAVM angiogenesis is characterized by loss of the intervening capillary bed and accumulation of arterial and venous ECs. Nidal tissue is characterized by high flow, immune cell recruitment, and expression of developmentally restricted angiogenic and BBB molecular programs. EC–mural cell crosstalk, unique inflammatory cell population recruitment, and endoMT characterize bAVM rupture states.
-
Models to elucidate bAVM pathogenesis and treatment paradigms
Many genetically engineered mouse and zebrafish models have been developed to elucidate syndromic (Table 3) and sporadic (Table 4) bAVM pathogenesis, each offering complementary strengths and limitations (97, 98). Syndromic (i.e., HHT and CM-AVM) models have been valuable in defining mechanisms underlying human disease genes, including TGF-β/BMP changes, and the interaction between inherited genetic factors and angiogenic stimuli. Consistent with the human disease, many syndromic models display systemic AVMs prone to rupture, confounding mortality and therapeutic success specific to bAVMs. In contrast, the zebrafish models depend on transient morpholinos and exhibit embryonic development of AVM-like lesions in embryonic vessel correlates that do not fully recapitulate the postnatal human disease (Table 3).
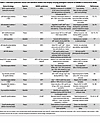 Table 3
Table 3Inherited syndromic mouse and zebrafish models that display varying pathogenic features of bAVMs or extracranial AVMs
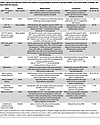 Table 4
Table 4Mouse and zebrafish models that display varying pathogenic features of sporadic bAVMs, extracranial AVMs, or bAVMs, and other AVM-like features
Since most human bAVMs are sporadic and driven by somatic KRAS — or less commonly BRAF — mutations, models expressing EC-specific human variants have provided critical mechanistic and therapeutic insights (Table 4). While no current model (adeno-associated virus–mediated [AAV-mediated] delivery or Cre-inducible transgenics) replicates the mosaicism and low pathogenic allele frequency seen in human sporadic bAVMs, they have been valuable in identifying a downstream dependency on MEK hyperactivation and for testing pharmacologic and gene editing strategies. However, these models likely overestimate disease penetrance, growth kinetics, and therapeutic response owing to the challenge of targeting rare cell populations, presence of approximately 1–3 bAVMs per mouse, and CV dysplasia beyond the bAVM in many cases. Nonetheless, EC-specific expression of KrasG12D or KrasG12V in mice results in multifocal extracranial and bAVMs (80, 99), and mice harboring brain-EC-specific expression of the driver mutation have been developed to overcome this limitation (100). Treatment of KrasG12D and KrasG12V mutant mice (Cre- or AAV-driven) with trametinib (MEK inhibitor) halted bAVM progression, reduced bAVM rupture, and normalized CV architecture (100–103). In addition, mice harboring EC-specific KrasG12C expression developed systemic and bAVMs that were ameliorated with sotorasib, a chemotherapeutic targeted therapy against the KrasG12C mutation (85). Overall, both syndromic and sporadic bAVM models provide a robust and reproducible means to define bAVM disease mechanisms and assess proof-of-principle therapeutic strategies.
-
Molecularly targeted therapies for bAVMs
Initial treatment approaches for bAVMs relied on antiangiogenic approaches (46, 104). However, with the discovery that somatic KRAS mutations cause most sporadic bAVMs, the potential of targeting KRAS activity has received special attention (105). Because KRAS driver mutations arise in ECs, therapeutic delivery — either small molecule or gene editing cargo — must be optimized for brain endothelial targeting. Notably, direct targeting of brain ECs does not require BBB penetration. The spatial resolution of mutant ECs to the arterial or venous side of the bAVM remains unresolved, but endovascular-enabled liquid biopsy of extracranial AVMs suggests that the draining vein may harbor the pathogenic variant (106). We hypothesize that endovascular-enabled therapeutic delivery (via transarterial or transvenous routes) may mitigate systemic pharmacologic toxicity challenges, and conjugation with liquid embolics could provide a long-term therapeutic depot. Development and/or modification of existing compounds to optimize mutation specificity, bAVM EC uptake, and long-term tolerability aimed at reprogramming bAVMs to a quiescent state (rather than complete volumetric obliteration) should be considered.
Given recent FDA recommendations on expediting testing and approval of cell- and gene-based therapies (107), precision medicine approaches for select, genetically defined bAVMs may be feasible. Nonetheless, selection of patient candidates for testing targeted therapy against bAVMs requires a multidisciplinary adjudication panel and detailed clinical monitoring. We believe that bAVM candidates for molecularly targeted therapy are those that are nonoperative (large or eloquent), actively growing, and/or radiation unresponsive. Importantly, pharmacotherapies targeting KRASG12C (sotorasib) and BRAFV600E (vemurafenib) are currently in clinical use for cancer and approved by the FDA. Sotorasib has produced marked, and in some cases, near-complete regression of KrasG12C bAVMs in mouse models and clinically meaningful regression of facial AVMs in humans (85), while vemurafenib has been shown to obliterate AVM-like lesions in zebrafish harboring BRAFV600E mutations (108). Additional targeted agents against KRASG12C (e.g., adagrasib, which has been shown to cross the BBB; ref. 109) and KRASG12D are available (110, 111), but require optimization for bAVM treatment. Trametinib has been used to treat one patient with a KRAS-mutant spinal AVM (112) and three pediatric bAVM patients (genotype unknown) with lesions deemed too large, in eloquent location, or untreatable by radiation (113). In each case, either growth or development of high-risk features prompted the experimental treatment (113). Two of three patients demonstrated stable disease while on therapy, while one patient’s bAVM hemorrhaged during trametinib treatment, prompting an increase in dose (1 mg/day from 0.5 mg/day) (113). However, the unknown lesional mutation status and short follow-up duration limits our understanding of how treatment might have changed the natural history of these bAVMs. Importantly, analysis of the response to therapy requires consensus on what defines “success” (e.g., cessation of bAVM growth, reversal of high-risk angioarchitecture, molecular response defined by reduction in variant allele fraction by liquid biopsy, etc.). Collectively, these data suggest that KRAS/BRAF mutation–specific or pathway-modulatory (i.e., MEK) pharmacotherapies are promising strategies for bAVM treatment.
-
Gene editing approaches for bAVMs
While molecularly targeted pharmacotherapies for bAVMs are appealing, gene editing technologies could, in principle, offer even greater precision, lasting effects, and a one-time genetic treatment (Figure 3). Limitations of CRISPR gene editing include editing specificity, targeting scope (determined by protospacer-adjacent motif [PAM] requirements), controlling and improving editing outcomes, and cell-targeted delivery methods (6, 7). Approaches to overcome specificity challenges include development of high-fidelity nuclease variants (114–116), modulating the duration of expression of the editor (117), and use of modified gRNAs (118, 119). The canonical 5′-NGG-3′ PAM sequence requirement of the Cas9 protein reduces targeting scope, but can be overcome using modified Cas9 or alternative Cas proteins with varying PAM requirements/specificities (120–130). Beyond traditional nuclease-based editing approaches, base editors can also introduce single nucleotide changes (131–134), and polymerization-based technologies, including prime editors (135) or DNA-dependent DNA polymerase–based editors (136, 137), can direct genomic DNA changes using RNA or DNA templates, respectively (Figure 3). These methods allow for more precise control of editing, particularly in the case of base or prime editing for single-point-mutation correction. Finally, additional gene editing tools include RNA editors (138) and epigenome editors (139, 140) that could be deployed in tandem with point-mutant-correction therapies (i.e., reduction of KRAS-dependent signaling) or as second-line therapies to modulate signaling pathways (i.e., VEGF, Notch, etc.) driving human bAVM maintenance, regression, response to radiation, and/or recurrence.
 Figure 3
Figure 3Gene editing strategies for bAVMs. (A) Strategies include CRISPR/Cas-based methods for gene knockout with nucleases to create insertions or deletions (indels) or templated edits via homology-directed repair. (B and C) Alternatively, more precise edits can be made with base editors and prime editors. (D) Delivery methods encompass lipid nanoparticles and (cerebro)vascular-specific AAVs, among others.
There are many potential challenges to gene editing for bAVMs. Specifically, human sporadic bAVMs display EC-restricted low variant allele fraction and mosaicism, requiring highly efficient and spatially restricted EC delivery. Recent advances in AAV engineering have expanded tropism to ECs in the cerebrovasculature (141–145), including a modified AAV-PR variant that transduces ECs and perivascular cells in murine models (144, 146). Lipid nanoparticles are an alternative approach, but important limitations include BBB permeability, where optimization of particle size, electrostatic properties, and selection of brain-EC-specific targeting ligands are potential clinical barriers (147–149). Nonetheless, in proof-of-concept experiments, CRISPR/Cas-mediated repression of KRAS expression has been shown to suppress bAVM growth and improve survival in KrasG12D mutant mice (99). Delivery method, immunogenicity, unintended nucleotide base conversions, introduction of genomic structural variations, and bystander edits stemming from nonspecific Cas9 binding or deaminase activity leading to unintended cellular perturbations remain key safety concerns for human clinical implementation. These data suggest that bAVMs may be suitable lesions for gene editing therapies, but a careful understanding of the disease biology, angioarchitectural markers of treatment success, and reliable detection and monitoring of the mutant EC fraction in situ will be necessary to guide the optimal gene editing approach, patient selection, and timing of treatment, among other factors.
-
In situ somatic mutation detection in CV pathology
The first and critical step towards implementing molecularly targeted therapies for bAVM treatment is detecting somatic driver mutations in situ. Endoluminal biopsy is one method to obtain CV lesional tissue (150, 151). In this approach, a microcatheter delivers a coil to the target vessel wall; after a brief dwell time, the coil is retrieved and adherent cells are isolated via flow cytometry (152, 153), typically focused on isolating ECs (154–156). Endoluminal biopsy has enabled capture of lesional tissue from an intracranial fusiform aneurysm to identify a somatic driver mutation in PDGFRB (157). Importantly, no complications from endoluminal biopsy have been reported (154–156). Endovascular-enabled liquid biopsy, on the other hand, captures cell-free DNA from peri-AVM or intranidal blood (106), which can be seamlessly integrated with existing endovascular clinical workflows. Important limitations that need further study include safety assessments, molecular sensitivity for detecting low-frequency mutations, and technical variability in somatic DNA yield. Developing a molecular diagnostic test that targets a few “actionable” mutations could facilitate genotype-guided treatment candidacy and molecular monitoring. Given the risks associated with bAVM rupture — and the understanding of the biological mechanisms involved — the potential risks of liquid and endoluminal biopsies may be justified in carefully selected patients with high-risk, otherwise untreatable lesions, although rigorous prospective clinical studies are necessary.
-
Integrated clinical-genomic framework for targeted therapy in bAVMs
How would precision medicine approaches be implemented in human patients? Rational selection of targeted therapies for bAVMs (deemed inoperable, untreatable with SRS, or persistent despite SRS) requires a minimally invasive method (i.e., liquid and/or endoluminal biopsy) for detecting somatic pathogenic variants. We suggest a precision medicine approach for bAVMs that includes (a) patient-parent trio whole-exome sequencing to detect germline mutations, (b) in situ bAVM somatic genotyping, and to guide (c) eligibility for mutation-specific or pathway modulatory agents or (e) bespoke gene editing strategies to correct pathogenic somatic mutations (Figure 4). However, consensus is needed on defining clinical, angiographic, and possibly molecular endpoints, patient selection criteria, and comparisons with natural history studies.
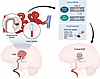 Figure 4
Figure 4Integrated clinical-genomic framework for precision medicine in bAVMs. Molecular diagnostics enabled by endoluminal and/or liquid biopsy for in situ detection of pathogenic somatic driver mutations to guide selection of molecularly targeted pharmacotherapy (sotorasib, vemurafenib, trametinib, etc.) or to identify pathogenic alleles for gene correction therapy in vivo (i.e., bespoke base editing approaches). Endoluminal and/or liquid biopsy can then be leveraged for postoperative bAVM surveillance in tandem with neuroimaging to monitor bAVM regression over time.
-
Conclusions
Rapid advances in human genetics, functional genomics, and CV biology have transformed our understanding of both sporadic and inherited bAVMs. Somatic activating mutations in KRAS — and, less frequently, BRAF — are now identified as common drivers in up to 80% of sporadic, surgically resected cases, while somatic second-hit mutations in ENG, ACVRL1, RASA1, and EPHB4 characterize inherited vascular syndromes variably linked to bAVMs. Alongside single-cell and spatial transcriptomic atlases and proof-of-principle animal studies, these findings create a mechanistic framework to consider precision therapy in human patients with bAVMs.
However, biological and translational gaps still exist. The timing, cadence, and hierarchy of bAVM development, genetic and environmental factors influencing growth and rupture, and the causal interaction among involved pathways are not fully understood. Many pathways implicated in model organisms often lack direct validation or relevance in human tissue, and phenotypic variability between lesions makes predicting therapeutic response more difficult. Early evidence for MEK- and KRAS-targeted treatments or CRISPR-based solutions is promising but still in the preclinical stage.
Emerging endovascular-enabled molecular diagnostics — liquid or endoluminal biopsy — may enable in situ detection of pathogenic mutations, although safety, reproducibility, and sensitivity need to be validated prospectively. Since current genomic analyses are almost entirely biased towards resected lesions, combining liquid or endovascular biopsy–enabled genotyping with angiographic anatomy could help prioritize subtypes for targeted intervention. Efforts towards clinical equipoise and multi-subspeciality coordination will be required for effective and safe translational efforts.
Ultimately, creation of an integrated genomic-phenomic taxonomy of bAVMs will redefine these malformations as treatable genetic diseases and guide genotype-directed therapeutic trials. Achieving this vision will require rigorous preclinical validation, cautious translation, and sustained collaboration across neurosurgery, interventional neuroradiology, vascular neurology, oncology, functional genomics, and bioengineering. The promise of precision medicine for bAVMs is real, but its implementation will demand both innovation and restraint.
-
Conflict of interest
ATH and KTK are co-inventors on a provisional patent application, 63/841,395, detailing molecular diagnostic approaches to enable precision medicine for bAVMs. Unrelated to this work, PKB has consulted for ElevateBio, Genentech, Angiochem, Tesaro, Axiom Healthcare Strategies, InCephalo Therapeutics, Medscape, MPM Capital Advisors, Dantari Pharmaceuticals, SK Life Sciences, Pfizer, CraniUS, Kazia, Sintetica, Voyager Therapeutics, Advise Connect Inspire, and Atavistik; served on the advisory board of CraniUS and Kazia; and has received research support (institution) from Merck, Mirati, Eli Lilly, and Kinnate. BPK is a consultant for Novartis Venture Fund, Generation Bio, and Jumble Therapeutics, and is on the scientific advisory boards of Life Edit Therapeutics and Prime Medicine. BPK has a financial interest in Prime Medicine, Inc., a company developing therapeutic CRISPR/Cas technologies for gene editing. All authors’ interests were reviewed and are managed by their respective institutions in accordance with their conflict-of-interest policies.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- NIH grants 1R01CA294793 and 5R01CA227156 (to PKB).
- NIH grants DP2CA281401 and P01HL142494 (to BPK).
- NIH grant R01NS125353 (to BPK and PLM).
- NIH grants R01NS111029, R01NS109358, and R01NS117609 (to KTK).
- National Institute on Aging grant F32AG089892 (to PQD).
- Aneurysm and AVM Foundation.
- Bee Foundation.
- Joe Niekro Foundation.
- American Association of Neurological Surgeons/Congress of Neurological Surgeons (AANS/CNS) Robert J. Dempsey, MD Research Award.
- Neurosurgery Research and Education Foundation (NREF).
- American Academy of Neurological Surgery (AAcNS).
- Society of Neurological Surgeons Neurosurgeon Scientist Training Program (to ATH).
- Breast Cancer Research Foundation (to PKB).
- National Brain Tumor Society (to PKB).
- Melanoma Research Alliance (to PKB).
- Demetra Fund from the Hellenic Women’s Club (to PKB).
- Terry and Jean de Gunzburg MGH Research Scholar Fund (to PKB).
- Kayden-Lambert MGH Research Scholar Award 2023-2028 (to BPK).
-
Acknowledgments
We apologize to colleagues whose studies we could not include due to space limitations. We would like to thank Laura Roy Illustrations for drafting Figure 4.
Address correspondence to: Andew T. Hale, 1720 2nd Ave. Birmingham, AL, 35233. Email: andrewthale20@gmail.com. Or to: Patricia L. Mulsolino, 55 Fruit St, Boston, MA 02114. Email: pmusolino@mgb.org. Or to: Kristopher T. Kahle, 15 Parkman St., WACC 331, Boston, MA 02114. Email: kahle.kristopher@mgh.harvard.edu.
-
Footnotes
Copyright: © 2026, Hale et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: J Clin Invest. 2026;136(10):e205086. https://doi.org/10.1172/JCI205086.
-
References
- Solomon RA, et al. Arteriovenous malformations of the brain. N Engl J Med. 2017;376(19):1859–1866.
- Berman MF, et al. The epidemiology of brain arteriovenous malformations. Neurosurgery. 2000;47(2):389–396; discussion 397.
- Rutledge WC, et al. Hemorrhage rates and risk factors in the natural history course of brain arteriovenous malformations. Transl Stroke Res. 2014;5(5):538–542.
- da Costa L, et al. The natural history and predictive features of hemorrhage from brain arteriovenous malformations. Stroke. 2009;40(1):100–105.
- Li H, et al. Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects. Signal Transduct Target Ther. 2020;5(1):1.
- Doudna JA. The promise and challenge of therapeutic genome editing. Nature. 2020;578(7794):229–236.
- Pacesa M, et al. Past, present, and future of CRISPR genome editing technologies. Cell. 2024;187(5):1076–1100.
- Goss JA, et al. Somatic mutations in intracranial arteriovenous malformations. PLoS One. 2019;14(12):e0226852.
- Hong T, et al. High prevalence of KRAS/BRAF somatic mutations in brain and spinal cord arteriovenous malformations. Brain. 2019;142(1):23–34.
- Nikolaev SI, et al. Somatic activating KRAS mutations in arteriovenous malformations of the brain. N Engl J Med. 2018;378(3):250–261.
- McDonald J, et al. Hereditary hemorrhagic telangiectasia: genetics and molecular diagnostics in a new era. Front Genet. 2015;6:1.
- McAllister KA, et al. Endoglin, a TGF-beta binding protein of endothelial cells, is the gene for hereditary haemorrhagic telangiectasia type 1. Nat Genet. 1994;8(4):345–351.
- Eerola I, et al. Capillary malformation-arteriovenous malformation, a new clinical and genetic disorder caused by RASA1 mutations. Am J Hum Genet. 2003;73(6):1240–1249.
- Zhao S, et al. Mutation of key signaling regulators of cerebrovascular development in vein of Galen malformations. Nat Commun. 2023;14(1):7452.
- Thomas-Sohl KA, et al. Sturge-Weber syndrome: a review. Pediatr Neurol. 2004;30(5):303–310.
- Carmeliet P, Jain RK. Molecular mechanisms and clinical applications of angiogenesis. Nature. 2011;473(7347):298–307.
- Abdelilah-Seyfried S, Ola R. Shear stress and pathophysiological PI3K involvement in vascular malformations. J Clin Invest. 2024;134(10):e172843.
- Tu T, et al. Neuroinflammation and hypoxia promote astrocyte phenotypic transformation and propel neurovascular dysfunction in brain arteriovenous malformation. J Neuroinflammation. 2025;22(1):124.
- Kiliç K, et al. Inhibition of angiogenesis induced by cerebral arteriovenous malformations using gamma knife irradiation. J Neurosurg. 2007;106(3):463–469.
- Hashimoto T, et al. Coexpression of angiogenic factors in brain arteriovenous malformations. Neurosurgery. 2005;56(5):1058–1065; discussion 1058.View this article via: PubMed Google Scholar
- Pawlikowska L, et al. Polymorphisms in genes involved in inflammatory and angiogenic pathways and the risk of hemorrhagic presentation of brain arteriovenous malformations. Stroke. 2004;35(10):2294–2300.
- Kuijper S, et al. Regulation of angiogenesis by Eph-ephrin interactions. Trends Cardiovasc Med. 2007;17(5):145–151.
- Zeng X, et al. EphrinB2-EphB4-RASA1 signaling in human cerebrovascular development and disease. Trends Mol Med. 2019;25(4):265–286.
- Wang HU, et al. Molecular distinction and angiogenic interaction between embryonic arteries and veins revealed by ephrin-B2 and its receptor Eph-B4. Cell. 1998;93(5):741–753.
- Bai J, et al. Ephrin B2 and EphB4 selectively mark arterial and venous vessels in cerebral arteriovenous malformation. J Int Med Res. 2014;42(2):405–415.
- Fehnel KP, et al. Dysregulation of the EphrinB2-EphB4 ratio in pediatric cerebral arteriovenous malformations is associated with endothelial cell dysfunction in vitro and functions as a novel noninvasive biomarker in patients. Exp Mol Med. 2020;52(4):658–671.
- Weinsheimer S, et al. EPHB4 gene polymorphisms and risk of intracranial hemorrhage in patients with brain arteriovenous malformations. Circ Cardiovasc Genet. 2009;2(5):476–482.
- Duran D, et al. Mutations in chromatin modifier and ephrin signaling genes in vein of galen malformation. Neuron. 2019;101(3):429–443.
- Vivanti A, et al. Loss of function mutations in EPHB4 are responsible for vein of Galen aneurysmal malformation. Brain. 2018;141(4):979–988.
- Vish KJ, et al. Diverse p120RasGAP interactions with doubly phosphorylated partners EphB4, p190RhoGAP, and Dok1. J Biol Chem. 2023;299(9):105098.
- Rak J, et al. Mutant ras oncogenes upregulate VEGF/VPF expression: implications for induction and inhibition of tumor angiogenesis. Cancer Res. 1995;55(20):4575–4580.View this article via: PubMed Google Scholar
- Li QF, et al. Activation of Ras in the vascular endothelium induces brain vascular malformations and hemorrhagic stroke. Cell Rep. 2018;24(11):2869–2882.
- Macmurdo CF, et al. RASA1 somatic mutation and variable expressivity in capillary malformation/arteriovenous malformation (CM/AVM) syndrome. Am J Med Genet A. 2016;170(6):1450–1454.
- Revencu N, et al. RASA1 mutations and associated phenotypes in 68 families with capillary malformation-arteriovenous malformation. Hum Mutat. 2013;34(12):1632–1641.
- Revencu N, et al. Parkes Weber syndrome, vein of Galen aneurysmal malformation, and other fast-flow vascular anomalies are caused by RASA1 mutations. Hum Mutat. 2008;29(7):959–965.
- Wooderchak-Donahue W, et al. RASA1 analysis: clinical and molecular findings in a series of consecutive cases. Eur J Med Genet. 2012;55(2):91–95.
- Chen D, et al. Angiogenesis depends upon EPHB4-mediated export of collagen IV from vascular endothelial cells. JCI Insight. 2022;7(4):e156928.
- Hatch AJ, York JD. SnapShot: Inositol phosphates. Cell. 2010;143(6):1030–1030.
- Castel P, et al. Somatic PIK3CA mutations as a driver of sporadic venous malformations. Sci Transl Med. 2016;8(332):332ra42.
- Peyre M, et al. Somatic PIK3CA mutations in sporadic cerebral cavernous malformations. N Engl J Med. 2021;385(11):996–1004.
- Kobialka P, et al. The onset of PI3K-related vascular malformations occurs during angiogenesis and is prevented by the AKT inhibitor miransertib. EMBO Mol Med. 2022;14(7):e15619.
- Iriarte A, et al. PI3K (phosphatidylinositol 3-kinase) activation and endothelial cell proliferation in patients with hemorrhagic hereditary telangiectasia type 1. Cells. 2019;8(9):971.
- Ola R, et al. PI3 kinase inhibition improves vascular malformations in mouse models of hereditary haemorrhagic telangiectasia. Nat Commun. 2016;7:13650.
- Nagy JA, et al. VEGF-A and the induction of pathological angiogenesis. Annu Rev Pathol. 2007;2:251–275.
- Lee CZ, et al. Matrix metalloproteinase-9 inhibition attenuates vascular endothelial growth factor-induced intracerebral hemorrhage. Stroke. 2007;38(9):2563–2568.
- Walker EJ, et al. Bevacizumab attenuates VEGF-induced angiogenesis and vascular malformations in the adult mouse brain. Stroke. 2012;43(7):1925–1930.
- Muster R, et al. Proof-of-concept single-arm trial of bevacizumab therapy for brain arteriovenous malformation. BMJ Neurol Open. 2021;3(1):e000114.
- Kamba T, McDonald DM. Mechanisms of adverse effects of anti-VEGF therapy for cancer. Br J Cancer. 2007;96(12):1788–1795.
- Wang K, et al. Perturbations of BMP/TGF-β and VEGF/VEGFR signalling pathways in non-syndromic sporadic brain arteriovenous malformations (BAVM). J Med Genet. 2018;55(10):675–684.
- dela Paz NG, et al. Role of shear-stress-induced VEGF expression in endothelial cell survival. J Cell Sci. 2012;125(pt 4):831–843.
- Buell TJ, et al. Embolization-induced angiogenesis in cerebral arteriovenous malformations. J Clin Neurosci. 2014;21(11):1866–1871.
- Zhang ZG, et al. VEGF enhances angiogenesis and promotes blood-brain barrier leakage in the ischemic brain. J Clin Invest. 2000;106(7):829–838.
- McDonald MT, et al. A disease locus for hereditary haemorrhagic telangiectasia maps to chromosome 9q33-34. Nat Genet. 1994;6(2):197–204.
- Lesca G, et al. Molecular screening of ALK1/ACVRL1 and ENG genes in hereditary hemorrhagic telangiectasia in France. Hum Mutat. 2004;23(4):289–299.
- Alsina-Sanchís E, et al. ALK1 loss results in vascular hyperplasia in mice and humans through PI3K activation. Arterioscler Thromb Vasc Biol. 2018;38(5):1216–1229.
- Chen W, et al. De novo cerebrovascular malformation in the adult mouse after endothelial Alk1 deletion and angiogenic stimulation. Stroke. 2014;45(3):900–902.
- Tual-Chalot S, et al. Endothelial depletion of Acvrl1 in mice leads to arteriovenous malformations associated with reduced endoglin expression. PLoS One. 2014;9(6):e98646.
- Zhu W, et al. Induction of brain arteriovenous malformation through CRISPR/Cas9-mediated somatic alk1 gene mutations in adult mice. Transl Stroke Res. 2019;10(5):557–565.
- Crist AM, et al. Vascular deficiency of Smad4 causes arteriovenous malformations: a mouse model of hereditary hemorrhagic telangiectasia. Angiogenesis. 2018;21(2):363–380.
- Crist AM, et al. Angiopoietin-2 inhibition rescues arteriovenous malformation in a Smad4 hereditary hemorrhagic telangiectasia mouse model. Circulation. 2019;139(17):2049–2063.
- Kim YH, et al. SMAD4 deficiency leads to development of arteriovenous malformations in neonatal and adult mice. J Am Heart Assoc. 2018;7(21):e009514.
- Ola R, et al. SMAD4 prevents flow induced arteriovenous malformations by inhibiting casein kinase 2. Circulation. 2018;138(21):2379–2394.
- Oppenheim O, et al. Divergent endothelial mechanisms drive arteriovenous malformations in Alk1 and SMAD4 loss-of-function [preprint]. https://doi.org/10.1101/2025.01.03.631070 Posted on bioRxiv January 3, 2025.
- Fu W, et al. Mesenchymal behavior of the endothelium promoted by SMAD6 downregulation is associated with brain arteriovenous malformation microhemorrhage. Stroke. 2020;51(7):2197–2207.
- Delev D, et al. NOTCH4 gene polymorphisms as potential risk factors for brain arteriovenous malformation development and hemorrhagic presentation. J Neurosurg. 2017;126(5):1552–1559.
- Kofler NM, et al. Combined deficiency of Notch1 and Notch3 causes pericyte dysfunction, models CADASIL, and results in arteriovenous malformations. Sci Rep. 2015;5:16449.
- Krebs LT, et al. Notch1 activation in mice causes arteriovenous malformations phenocopied by ephrinB2 and EphB4 mutants. Genesis. 2010;48(3):146–150.
- Murphy PA, et al. Constitutively active Notch4 receptor elicits brain arteriovenous malformations through enlargement of capillary-like vessels. Proc Natl Acad Sci U S A. 2014;111(50):18007–18012.
- Murphy PA, et al. Endothelial Notch4 signaling induces hallmarks of brain arteriovenous malformations in mice. Proc Natl Acad Sci U S A. 2008;105(31):10901–10906.
- ZhuGe Q, et al. Notch-1 signalling is activated in brain arteriovenous malformations in humans. Brain. 2009;132(pt 12):3231–3241.
- Diéguez-Hurtado R, et al. Loss of the transcription factor RBPJ induces disease-promoting properties in brain pericytes. Nat Commun. 2019;10(1):2817.
- Nielsen CM, et al. Deletion of Rbpj from postnatal endothelium leads to abnormal arteriovenous shunting in mice. Development. 2014;141(19):3782–3792.
- Nielsen CM, et al. Endothelial Rbpj deletion normalizes Notch4-induced brain arteriovenous malformation in mice. J Exp Med. 2023;220(2):e20211390.
- Krebs LT, et al. Haploinsufficient lethality and formation of arteriovenous malformations in Notch pathway mutants. Genes Dev. 2004;18(20):2469–2473.
- Nadeem T, et al. Deficiency of Notch signaling in pericytes results in arteriovenous malformations. JCI Insight. 2020;5(21):e125940.
- Karthika CL, et al. Oscillatory shear stress modulates Notch-mediated endothelial mesenchymal plasticity in cerebral arteriovenous malformations. Cell Mol Biol Lett. 2023;28(1):22.
- Huang L, et al. Nitric oxide synthase and reduced arterial tone contribute to arteriovenous malformation. Sci Adv. 2023;9(21):eade7280.
- Tu J, et al. Radiosurgery inhibition of the Notch signaling pathway in a rat model of arteriovenous malformations. J Neurosurg. 2014;120(6):1385–1396.
- Xu H, et al. KRAS mutation-induced EndMT of brain arteriovenous malformation is mediated through the TGF-β/BMP-SMAD4 pathway. Stroke Vasc Neurol. 2023;8(3):197–206.
- Fish JE, et al. Somatic gain of KRAS function in the endothelium is sufficient to cause vascular malformations that require MEK but not PI3K signaling. Circ Res. 2020;127(6):727–743.
- Amyere M, et al. Germline loss-of-function mutations in EPHB4 cause a second form of capillary malformation-arteriovenous malformation (CM-AVM2) deregulating RAS-MAPK signaling. Circulation. 2017;136(11):1037–1048.
- Shirley MD, et al. Sturge-Weber syndrome and port-wine stains caused by somatic mutation in GNAQ. N Engl J Med. 2013;368(21):1971–1979.
- Bernabeu C, et al. Potential second-hits in hereditary hemorrhagic telangiectasia. J Clin Med. 2020;9(11):3571.
- Larrivée B, et al. ALK1 signaling inhibits angiogenesis by cooperating with the Notch pathway. Dev Cell. 2012;22(3):489–500.
- Fraissenon A, et al. Sotorasib for vascular malformations associated with KRAS G12C mutation. N Engl J Med. 2024;391(4):334–342.
- Mikhak B, et al. Angiopoietin-like 4 (ANGPTL4) gene polymorphisms and risk of brain arteriovenous malformations. Cerebrovasc Dis. 2011;31(4):338–345.
- Li H, et al. De novo germline and somatic variants convergently promote endothelial-to-mesenchymal transition in simplex brain arteriovenous malformation. Circ Res. 2021;129(9):825–839.
- Wang K, et al. Exome-wide analysis of de novo and rare genetic variants in patients with brain arteriovenous malformation. Neurology. 2022;98(16):1670–1678.
- Zhang M, et al. Exome sequencing of 112 trios identifies recessive genetic variants in brain arteriovenous malformations. J Neurointerv Surg. 2021;13(6):568–573.
- Crouch EE, et al. Ensembles of endothelial and mural cells promote angiogenesis in prenatal human brain. Cell. 2022;185(20):3753–3769.
- Garcia FJ, et al. Single-cell dissection of the human brain vasculature. Nature. 2022;603(7903):893–899.
- Wälchli T, et al. Single-cell atlas of the human brain vasculature across development, adulthood and disease. Nature. 2024;632(8025):603–613.
- Winkler EA, et al. A single-cell atlas of the normal and malformed human brain vasculature. Science. 2022;375(6584):eabi7377.
- Lippmann ES, et al. Derivation of blood-brain barrier endothelial cells from human pluripotent stem cells. Nat Biotechnol. 2012;30(8):783–791.
- Li R, et al. Androgen receptor governs tip cell formation in cerebrovascular malformations. Circ Res. 2025;137(5):628–645.
- Scimone C, et al. Methylome analysis of endothelial cells suggests new insights on sporadic brain arteriovenous malformation. Heliyon. 2024;10(15):e35126.
- Nielsen CM, et al. Mouse models of cerebral arteriovenous malformation. Stroke. 2016;47(1):293–300.
- Ricciardelli AR, et al. From bench to bedside: murine models of inherited and sporadic brain arteriovenous malformations. Angiogenesis. 2025;28(2):15.
- Saito S, et al. CRISPR/CasRx suppresses KRAS-induced brain arteriovenous malformation developed in postnatal brain endothelial cells in mice. JCI Insight. 2024;9(22):e179729.
- Suarez CPF, et al. MEK signaling represents a viable therapeutic vulnerability of KRAS-driven somatic brain arteriovenous malformations [preprint]. https://doi.org/10.1101/2024.05.15.594335 Posted on bioRxiv May 15, 2024.
- Nguyen HL, et al. Trametinib as a promising therapeutic option in alleviating vascular defects in an endothelial KRAS-induced mouse model. Hum Mol Genet. 2023;32(2):276–289.
- Park ES, et al. Selective endothelial hyperactivation of oncogenic KRAS induces brain arteriovenous malformations in mice. Ann Neurol. 2021;89(5):926–941.
- Myint O, et al. Trametinib decreased intracerebral hemorrhages and endothelial-to-mesenchymal transition in KRASG12V-induced brain arteriovenous malformations in mice. Stroke. 2025;57(3):779–793.
- Zhu W, et al. Thalidomide reduces hemorrhage of brain arteriovenous malformations in a mouse model. Stroke. 2018;49(5):1232–1240.
- Hale AT, et al. An opportune time for targeted brain arteriovenous malformation therapy. J Neurosurg. 2025;143(6):1685–1688.
- Bhatia KD, et al. Harnessing liquid biopsy to unveil RAS-MEK pathway somatic pathogenic variants in extracranial arterio-venous malformations. Commun Med (Lond). 2025;5(1):508.
- Prasad V, Makary MA. FDA’s new plausible mechanism pathway. N Engl J Med. 2025;393(23):2365–2367.
- Al-Olabi L, et al. Mosaic RAS/MAPK variants cause sporadic vascular malformations which respond to targeted therapy. J Clin Invest. 2018;128(4):1496–1508.
- Sabari JK, et al. Activity of adagrasib (MRTX849) in brain metastases: preclinical models and clinical data from patients with KRASG12C-mutant non-small cell lung cancer. Clin Cancer Res. 2022;28(15):3318–3328.
- Hallin J, et al. Anti-tumor efficacy of a potent and selective non-covalent KRASG12D inhibitor. Nat Med. 2022;28(10):2171–2182.
- Jänne PA, et al. Adagrasib in non-small-cell lung cancer harboring a KRASG12C mutation. N Engl J Med. 2022;387(2):120–131.
- Cooke DL, et al. Angiographic evidence of response to trametinib therapy for a spinal cord arteriovenous malformation. J Vasc Anom. 2021;2(3):e018.
- Suarez CF, et al. MEK signaling represents a viable therapeutic vulnerability of KRAS-driven somatic brain arteriovenous malformations [preprint]. https://doi.org/10.1101/2024.05.15.594335 Posted on bioRxiv May 15, 2024.
- Chen JS, et al. Enhanced proofreading governs CRISPR-Cas9 targeting accuracy. Nature. 2017;550(7676):407–410.
- Kleinstiver BP, et al. High-fidelity CRISPR-Cas9 nucleases with no detectable genome-wide off-target effects. Nature. 2016;529(7587):490–495.
- Slaymaker IM, et al. Rationally engineered Cas9 nucleases with improved specificity. Science. 2016;351(6268):84–88.
- Kim S, et al. Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res. 2014;24(6):1012–1019.
- Donohoue PD, et al. Conformational control of Cas9 by CRISPR hybrid RNA-DNA guides mitigates off-target activity in T cells. Mol Cell. 2021;81(17):3637–3649.
- Hendel A, et al. Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat Biotechnol. 2015;33(9):985–989.
- Chatterjee P, et al. An engineered ScCas9 with broad PAM range and high specificity and activity. Nat Biotechnol. 2020;38(10):1154–1158.
- Esvelt KM, et al. Orthogonal Cas9 proteins for RNA-guided gene regulation and editing. Nat Methods. 2013;10(11):1116–1121.
- Gao L, et al. Engineered Cpf1 variants with altered PAM specificities. Nat Biotechnol. 2017;35(8):789–792.
- Kleinstiver BP, et al. Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat Biotechnol. 2015;33(12):1293–1298.
- Kleinstiver BP, et al. Engineered CRISPR-Cas9 nucleases with altered PAM specificities. Nature. 2015;523(7561):481–485.
- Kleinstiver BP, et al. Engineered CRISPR-Cas12a variants with increased activities and improved targeting ranges for gene, epigenetic and base editing. Nat Biotechnol. 2019;37(3):276–282.
- Miller SM, et al. Continuous evolution of SpCas9 variants compatible with non-G PAMs. Nat Biotechnol. 2020;38(4):471–481.
- Nishimasu H, et al. Engineered CRISPR-Cas9 nuclease with expanded targeting space. Science. 2018;361(6408):1259–1262.
- Silverstein RA, et al. Custom CRISPR-Cas9 PAM variants via scalable engineering and machine learning. Nature. 2025;643(8071):539–550.
- Walton RT, et al. Unconstrained genome targeting with near-PAMless engineered CRISPR-Cas9 variants. Science. 2020;368(6488):290–296.
- Zetsche B, et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell. 2015;163(3):759–771.
- Gaudelli NM, et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature. 2017;551(7681):464–471.
- Komor AC, et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature. 2016;533(7603):420–424.
- Nishida K, et al. Targeted nucleotide editing using hybrid prokaryotic and vertebrate adaptive immune systems. Science. 2016;353(6305):aaf8729.
- Porto EM, Komor AC. In the business of base editors: evolution from bench to bedside. PLoS Biol. 2023;21(4):e3002071.
- Anzalone AV, et al. Search-and-replace genome editing without double-strand breaks or donor DNA. Nature. 2019;576(7785):149–157.
- Ferreira da Silva J, et al. Click editing enables programmable genome writing using DNA polymerases and HUH endonucleases. Nat Biotechnol. 2024;43(6):923–935.
- Liu B, et al. Targeted genome editing with a DNA-dependent DNA polymerase and exogenous DNA-containing templates. Nat Biotechnol. 2024;42(7):1039–1045.
- Cox DBT, et al. RNA editing with CRISPR-Cas13. Science. 2017;358(6366):1019–1027.
- Kungulovski G, Jeltsch A. Epigenome editing: state of the art, concepts, and perspectives. Trends Genet. 2016;32(2):101–113.
- McCutcheon SR, et al. Epigenome editing technologies for discovery and medicine. Nat Biotechnol. 2024;42(8):1199–1217.
- Körbelin J, et al. A brain microvasculature endothelial cell-specific viral vector with the potential to treat neurovascular and neurological diseases. EMBO Mol Med. 2016;8(6):609–625.
- Krolak T, et al. A high-efficiency AAV for endothelial cell transduction throughout the central nervous system. Nat Cardiovasc Res. 2022;1(4):389–400.
- Li JL, et al. A targeted vector for brain endothelial cell gene delivery and cerebrovascular malformation modelling. Nat Biomed Eng. 2025;
- Ramirez SH, et al. An engineered adeno-associated virus capsid mediates efficient transduction of pericytes and smooth muscle cells of the brain vasculature. Hum Gene Ther. 2023;34(15-16):682–696.
- Velazquez-Rivera E, et al. Specific targeting of brain endothelial cells using enhancer AAV vectors. Neuron. 2025;113(10):1562–1578.
- Alves CRR, et al. Treatment of a severe vascular disease using a bespoke CRISPR-Cas9 base editor in mice. Nat Biomed Eng. 2025;
- Blasi P, et al. Solid lipid nanoparticles for targeted brain drug delivery. Adv Drug Deliv Rev. 2007;59(6):454–477.
- Hou X, et al. Lipid nanoparticles for mRNA delivery. Nat Rev Mater. 2021;6(12):1078–1094.
- Pardridge WM. Brain gene therapy with Trojan horse lipid nanoparticles. Trends Mol Med. 2023;29(5):343–353.
- Winkler EA, et al. Endoluminal biopsy for molecular profiling of human brain vascular malformations. Neurology. 2022;98(16):1637–1647.
- Cooke DL, et al. Endovascular biopsy: technical feasibility of novel endothelial cell harvesting devices assessed in a rabbit aneurysm model. Interv Neuroradiol. 2015;21(1):120–128.
- Alexander MD, et al. Assessment of cell yield among different devices for endovascular biopsy to harvest vascular endothelial cells. Biotechniques. 2019;66(1):34–36.
- Sun Z, et al. Endothelial cell high-enrichment from endovascular biopsy sample by laser capture microdissection and fluorescence activated cell sorting. J Biotechnol. 2014;192 Pt A(0 0):34–39.
- Hale AT, et al. Endoluminal biopsy for vein of galen malformation. Neurosurgery. 2024;95(5):1082–1088.
- Winkler E, et al. Endoluminal biopsy for molecular profiling of human brain vascular malformations. Neurology. 2022;98(16):1637–1647.
- Cooke DL, et al. Endovascular biopsy: in vivo cerebral aneurysm endothelial cell sampling and gene expression analysis. Transl Stroke Res. 2018;9(1):20–33.
- Garcia JH, et al. Endovascular biopsy for detection of somatic mosaicism in human fusiform cerebral aneurysms. Stroke Vasc Interv Neurol. 2022;2(5):e000354.
- Lesca G, et al. Distribution of ENG and ACVRL1 (ALK1) mutations in French HHT patients. Hum Mutat. 2006;27(6):598.
- Letteboer TG, et al. Hereditary hemorrhagic telangiectasia: ENG and ALK-1 mutations in Dutch patients. Hum Genet. 2005;116(1-2):8–16.
- Snellings DA, et al. Somatic mutations in vascular malformations of hereditary hemorrhagic telangiectasia result in bi-allelic loss of ENG or ACVRL1. Am J Hum Genet. 2019;105(5):894–906.
- Wooderchak-Donahue WL, et al. BMP9 mutations cause a vascular-anomaly syndrome with phenotypic overlap with hereditary hemorrhagic telangiectasia. Am J Hum Genet. 2013;93(3):530–537.
- Nakashima M, et al. The somatic GNAQ mutation c.548G>A (p.R183Q) is consistently found in Sturge-Weber syndrome. J Hum Genet. 2014;59(12):691–693.
- Gallione CJ, et al. SMAD4 mutations found in unselected HHT patients. J Med Genet. 2006;43(10):793–797.
- Oka M, et al. KRAS G12D or G12V mutation in human brain arteriovenous malformations. World Neurosurg. 2019;126:1365–1373.
- Priemer DS, et al. Activating KRAS mutations in arteriovenous malformations of the brain: frequency and clinicopathologic correlation. Hum Pathol. 2019;89:33–39.
- Gao S, et al. Somatic mosaicism in the MAPK pathway in sporadic brain arteriovenous malformation and association with phenotype. J Neurosurg. 2022;136(1):148–155.
- Erkinova SA, et al. Angiopoietin-like proteins 4 (ANGPTL4) gene polymorphisms and risk of brain arteriovenous malformation. J Stroke Cerebrovasc Dis. 2018;27(4):908–913.
- Walcott BP, et al. Identification of a rare BMP pathway mutation in a non-syndromic human brain arteriovenous malformation via exome sequencing. Hum Genome Var. 2018;5:18001.
- Scimone C, et al. High-throughput sequencing to detect novel likely gene-disrupting variants in pathogenesis of sporadic brain arteriovenous malformations. Front Genet. 2020;11:146.
- Park SO, et al. ALK5- and TGFBR2-independent role of ALK1 in the pathogenesis of hereditary hemorrhagic telangiectasia type 2. Blood. 2008;111(2):633–642.
- Park SO, et al. Real-time imaging of de novo arteriovenous malformation in a mouse model of hereditary hemorrhagic telangiectasia. J Clin Invest. 2009;119(11):3487–3496.
- Srinivasan S, et al. A mouse model for hereditary hemorrhagic telangiectasia (HHT) type 2. Hum Mol Genet. 2003;12(5):473–482.
- Hao Q, et al. VEGF induces more severe cerebrovascular dysplasia in endoglin than in Alk1 mice. Transl Stroke Res. 2010;1(3):197–201.
- Choi EJ, et al. Minimal homozygous endothelial deletion of Eng with VEGF stimulation is sufficient to cause cerebrovascular dysplasia in the adult mouse. Cerebrovasc Dis. 2012;33(6):540–547.
- Walker EJ, et al. Arteriovenous malformation in the adult mouse brain resembling the human disease. Ann Neurol. 2011;69(6):954–962.
- Chen W, et al. Reduced mural cell coverage and impaired vessel integrity after angiogenic stimulation in the Alk1-deficient brain. Arterioscler Thromb Vasc Biol. 2013;33(2):305–310.
- Corti P, et al. Interaction between alk1 and blood flow in the development of arteriovenous malformations. Development. 2011;138(8):1573–1582.
- Milton I, et al. Age-dependent lethality in novel transgenic mouse models of central nervous system arteriovenous malformations. Stroke. 2012;43(5):1432–1435.
- Han C, et al. Novel experimental model of brain arteriovenous malformations using conditional Alk1 gene deletion in transgenic mice. J Neurosurg. 2022;137(1):163–174.
- Choi H, et al. BMP10 functions independently from BMP9 for the development of a proper arteriovenous network. Angiogenesis. 2023;26(1):167–186.
- Bourdeau A, et al. A murine model of hereditary hemorrhagic telangiectasia. J Clin Invest. 1999;104(10):1343–1351.
- Torsney E, et al. Mouse model for hereditary hemorrhagic telangiectasia has a generalized vascular abnormality. Circulation. 2003;107(12):1653–1657.
- Satomi J, et al. Cerebral vascular abnormalities in a murine model of hereditary hemorrhagic telangiectasia. Stroke. 2003;34(3):783–789.
- Jin Y, et al. Endoglin prevents vascular malformation by regulating flow-induced cell migration and specification through VEGFR2 signalling. Nat Cell Biol. 2017;19(6):639–652.
- Xu B, et al. Vascular endothelial growth factor induces abnormal microvasculature in the endoglin heterozygous mouse brain. J Cereb Blood Flow Metab. 2004;24(2):237–244.
- Choi EJ, et al. Novel brain arteriovenous malformation mouse models for type 1 hereditary hemorrhagic telangiectasia. PLoS One. 2014;9(2):e88511.
- Zhou X, et al. ANG2 blockade diminishes proangiogenic cerebrovascular defects associated with models of hereditary hemorrhagic telangiectasia. Arterioscler Thromb Vasc Biol. 2023;43(8):1384–1403.
- Li JL, et al. rAAV-miniBEND: A targeted vector for brain endothelial cell gene delivery and cerebrovascular malformation modeling [published online October 29, 2025]. Nat Biomed Eng. https://doi.org/10.1038/s41551-025-01538-x.
- Tu T, et al. Somatic BrafV600E mutation in the cerebral endothelium induces brain arteriovenous malformations. Angiogenesis. 2024;27(3):441–460.
- Smits PJ, et al. Endothelial cell expression of mutant Map2k1 causes vascular malformations in mice. Angiogenesis. 2023;26(1):97–105.
- Yao Y, et al. Reducing Jagged 1 and 2 levels prevents cerebral arteriovenous malformations in matrix Gla protein deficiency. Proc Natl Acad Sci U S A. 2013;110(47):19071–19076.
- Carlson TR, et al. Endothelial expression of constitutively active Notch4 elicits reversible arteriovenous malformations in adult mice. Proc Natl Acad Sci U S A. 2005;102(28):9884–9889.
- Murphy PA, et al. Notch4 normalization reduces blood vessel size in arteriovenous malformations. Sci Transl Med. 2012;4(117):117ra8.
- Selhorst S, et al. Pathological pericyte expansion and impaired endothelial cell-pericyte communication in endothelial Rbpj deficient brain arteriovenous malformation. Front Hum Neurosci. 2022;16:974033.
-
Version history
- Version 1 (May 15, 2026): Electronic publication
Article tools
-
View PDF

- Download citation information
- Send a comment
- Terms of use
- Standard abbreviations
- Need help? Email the journal
Metrics
Go to
- Top
- Abstract
- Introduction
- Clinical overview of bAVMs
- Angiogenesis, AV specification, zonation, and CV development
- Genetic basis of bAVM pathogenesis
- Single-cell analysis of the cerebrovasculature and bAVMs
- Models to elucidate bAVM pathogenesis and treatment paradigms
- Molecularly targeted therapies for bAVMs
- Gene editing approaches for bAVMs
- In situ somatic mutation detection in CV pathology
- Integrated clinical-genomic framework for targeted therapy in bAVMs
- Conclusions
- Conflict of interest
- Funding support
- Acknowledgments
- Footnotes
- References
- Version history



Copyright © 2026 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)








