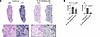Advertisement
Research ArticleImmunologyPulmonology
Open Access |  10.1172/JCI188891
10.1172/JCI188891
Impaired complement regulation drives chronic lung allograft dysfunction after lung transplantation
Hrishikesh S. Kulkarni,1,2 Laneshia K. Tague,1 Daniel R. Calabrese,3,4 Fuyi Liao,5 Zhiyi Liu,5 Lorena Garnica,1 Nishanth R. Shankar,1 Xiaobo Wu,1 Devesha H. Kulkarni,2 Aayusha Thapa,2 Dequan Zhou,5 Yan Tao,1 Victoria E. Davis,5 Cory T. Bernardt,6 Derek E. Byers,1 Catherine Chen,7 Howard J. Huang,8 Chad A. Witt,1 Ramsey R. Hachem,9 Daniel Kreisel,5,6 John P. Atkinson,1 John R. Greenland,3,4 and Andrew E. Gelman5,6
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by
Kulkarni, H.
in:
PubMed
|
Google Scholar
|

1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by
Tague, L.
in:
PubMed
|
Google Scholar
|

1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by
Calabrese, D.
in:
PubMed
|
Google Scholar
|

1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Liao, F. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Liu, Z. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Garnica, L. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Shankar, N. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Wu, X. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Kulkarni, D. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Thapa, A. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Zhou, D. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Tao, Y. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Davis, V. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Bernardt, C. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by
Byers, D.
in:
PubMed
|
Google Scholar
|

1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Chen, C. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by
Huang, H.
in:
PubMed
|
Google Scholar
|

1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Witt, C. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Hachem, R. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Kreisel, D. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by Atkinson, J. in: PubMed | Google Scholar
1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by
Greenland, J.
in:
PubMed
|
Google Scholar
|

1Department of Medicine, Washington University School of Medicine, St. Louis, Missouri, USA.
2Department of Medicine, David Geffen School of Medicine at UCLA, Los Angeles, CA, USA.
3Department of Medicine, UCSF, San Francisco, California, USA.
4Medical Service, Veterans Affairs Health Care System, San Francisco, California, USA.
5Department of Surgery and
6Department of Pathology & Immunology, Washington University School of Medicine, St. Louis, Missouri, USA.
7Department of Internal Medicine, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
8Department of Medicine, Houston Methodist Hospital, Houston, Texas, USA.
9Department of Internal Medicine, University of Utah Spencer Fox Eccles School of Medicine, Salt Lake City, Utah, USA.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
Find articles by
Gelman, A.
in:
PubMed
|
Google Scholar
|

Published November 11, 2025 - More info
J Clin Invest. 2026;136(1):e188891. https://doi.org/10.1172/JCI188891.
© 2025 Kulkarni et al. This work is licensed under the Creative Commons Attribution 4.0 International License. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
Received: November 18, 2024; Accepted: November 6, 2025
-
Abstract
A greater understanding of chronic lung allograft dysfunction (CLAD) pathobiology, the primary cause of death after lung transplantation (LTx), is needed to improve outcomes. The complement system links innate to adaptive immune responses and is activated early after lung transplantation to form C3 convertase, a critical enzyme that cleaves the central complement component C3. We hypothesized that LTx recipients with a genetic predisposition to enhanced complement activation have worse CLAD-free survival mediated through increased adaptive alloimmunity. We interrogated a known functional C3 polymorphism (C3 R102G) that increases complement activation through impaired C3 convertase inactivation in 2 independent LTx recipient cohorts. C3 R102G, identified in at least 1 of 3 LTx recipients, was associated with worse CLAD-free survival, particularly in the subset of recipients who developed donor-specific antibodies (DSAs). In a mouse orthotopic LTx model, impaired recipient complement regulation led to B cell–dependent CLAD pathology despite moderate differences in graft-infiltrating effector T cells. Dysfunctional complement regulation promoted intragraft accumulation of memory B cells and Ab-secreting cells, leading to increased local and circulating DSA levels in mice. In summary, genetic predisposition to complement activation is associated with an increased humoral response and worse CLAD-free survival.
-
Introduction
Chronic lung allograft dysfunction (CLAD) is the primary cause of long-term morbidity and mortality after lung transplantation (LTx) (1, 2). CLAD commonly presents with a constrictive bronchiolitis pathology and is associated with poor survival (3). However, it is unclear why some LTx recipients progress to CLAD whereas others do not. CLAD is also a heterogenous syndrome with greater or lesser involvement of donor-specific antibodies (DSAs) and alveolar space, leading to Ab-mediated rejection (AMR) and restrictive allograft syndrome (RAS) (4–6). Crosstalk between innate and adaptive immunity via the complement system may be necessary for driving CLAD (7–9). Although T cells are required for acute and chronic rejection, there is increasing evidence that B cells contribute to poor LTx survival, potentially through AMR (10–12). Additionally, mouse models of LTx have demonstrated B cell–dependent CLAD-like pathology (13–15).
The complement system comprises more than 50 proteins that are activated, amplified, and deposited onto pathogens and dying cells to facilitate their clearance by phagocytosis (16). This system can also be activated in transplantation via Ab-dependent and Ab-independent pathways (17–20). C3a and C5a are cleaved from C3 and C5 upon activating this cascade and can engage with their cognate receptors to influence alloimmune responses (21). Engagement of these receptors also drives profibrotic responses (22, 23). At the same time, the key complement protein C3 is required for the survival of both immune (24) and nonimmune (25–27) cells and modulates effector cell function (28).
A series of both membrane-bound and fluid-phase regulators controls complement activation and facilitates tissue homeostasis (29). Genetic and acquired deficiencies in these regulators increase complement activation, resulting in augmented tissue damage (30, 31). CD46, a membrane regulator, is a cofactor for fluid-phase regulators such as Factor I to cleave C3b and C4b and inactivates C3 convertase to attenuate complement activation. Crry, a murine ortholog of CD46, prevents complement activation on the vascular endothelium (32). Crry is downregulated during lung injury (33) and has also been used to ameliorate lung injury (34, 35).
The role of complement regulation in CLAD remains undefined. Given the centrality of C3 in the cascade, we analyzed a functional polymorphism in C3 (rs2231099, C3 R102G) that has a known minor allele frequency of at least 10% in several large cohorts. This polymorphism results in increased complement activation due to the reduced ability of the fluid-phase regulator Factor H to inactivate C3 convertase (36). Previous work has shown that the C3 R102G polymorphism is linked to the development of age-related macular degeneration and other complement-related conditions (37). We hypothesized that C3 R102G is associated with CLAD development. We show that recipients carrying the C3 R102G polymorphism in 2 independent human cohorts had worse CLAD-free survival, and these outcomes were more pronounced in the setting of DSAs. Additionally, using an orthotopic mouse LTx model, we demonstrate that recipients deficient in the complement regulator Crry developed CLAD caused by elevated intragraft B cell activation and high DSA levels (38, 39).
-
Results
A functional polymorphism in the complement component C3 gene is observed in at least 1 in 3 LTx recipients. Within the LTx cohort at Washington University/Barnes-Jewish Hospital (BJH), we identified 392 participants with available genotyping (Figure 1A). Among these recipients, 244 (62.2%) had wild-type C3 (G/G); thus, the recipient rs2230199 (G>C) SNP had a minor allele frequency (allele C) of 37.8%. Baseline characteristics for participants stratified by genotype are shown in Table 1 and were similar across both genotypes.
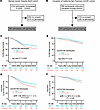 Figure 1
Figure 1A functional C3 polymorphism confers increased risk of CLAD or death in 2 independent cohorts. Consolidated Standards of Reporting Trials diagrams for the Washington University/BJH (A) and UCSF cohorts (B). (C and E) Kaplan-Meier plot of survival from CLAD or death in the BJH (C) and UCSF (E) cohorts stratified by rs2230199 genotypes. (D and F) Kaplan-Meier plot based on the multivariate Cox regression analysis of CLAD-free survival in the BJH (D) and UCSF (F) cohorts by rs2230199 status. The P value was determined by log-rank test.
Within the UCSF LTx cohort, we identified 425 participants with available genotyping (Figure 1B). The recipient rs2230199 (G>C) SNP had a minor allele frequency (allele C) of 29.4%. Baseline characteristics for participants stratified by genotype are shown in Table 2. Notably, we found higher minor allele frequencies among White recipients and lower minor allele frequencies among Black recipients. These distributions are consistent with those reported in aggregated global genomic databases (40). There were no other differences in LTx baseline characteristics across genotypes.
C3 R102G confers increased risk of CLAD or death in 2 independent cohorts. We next examined whether the rs2230199 C3 minor allele, which predisposes to increased complement activation, was associated with decreased CLAD-free survival. In the BJH cohort, the C3 R102G polymorphism was associated with an increased risk for CLAD or death (Figure 1C). In a multivariable Cox-proportional hazards model consisting of C3 rs2230199 genotype, age, sex, race, transplant diagnosis, DSAs, and Gram-negative infection, C3 R102G remained associated with increased risk of CLAD or death (Figure 1D and Table 3). The presence of post-transplant DSAs (adjusted hazard ratio [aHR] 1.74; 95% CI, 1.24–2.45; P = 0.001) or post-transplant Gram-negative infection (aHR 1.44; 95% CI, 1.01–2.06; P = 0.044) also was associated with a significantly increased risk of CLAD or death. A sensitivity analysis removing the presence of DSAs among the list of variables demonstrated that C3 R102G was still associated with an increased risk of CLAD or death (aHR 1.52; 95% CI, 1.08–2.15; P = 0.016). A similar analysis removing the presence of Gram-negative infection among the list of variables demonstrated that C3 R102G remained associated with an increased risk of CLAD or death (aHR 1.43; 95% CI, 1.02–2.02; P = 0.04).
We examined if R102G also was associated with differential CLAD-free survival in the UCSF cohort. Figure 1E shows that participants homozygous for C3 R102G had an increased risk for CLAD or death (HR 1.43; 95% CI, 1–2.05; adjusted P = 0.048). This multivariable Cox-proportional hazards model also consisted of C3 rs2230199 genotype, age, sex, race, transplant diagnosis, DSAs, and Gram-negative infection (Figure 1F and Table 4).
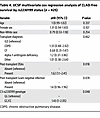 Table 4
Table 4UCSF multivariate cox regression analysis of CLAD-free survival by rs2230199 status (n = 425)
CLAD-free survival association with recipient C3 R102G is linked to DSAs. Based on work published by us and others reporting that anti-HLA DSAs are a known risk factor for CLAD (11, 41), we hypothesized that the increased risk of CLAD or death in recipients with C3 R102G would be limited to those with DSAs. In the BJH cohort, 181 LTx recipients (46.2%) had definite DSAs during their follow-up. CLAD-free survival analysis accounting for both DSAs and rs2230199 genotype demonstrated that recipients with C3 R102G and DSAs had the worst CLAD-free survival (Figure 2A). Among recipients without DSAs, we did not observe a difference in risk for CLAD or death based on the C3 rs2230199 genotype (HR 1.21; 95% CI, 0.72–2.04; P = 0.48). However, among the participants who developed DSAs and had the C3 R102G polymorphism, there was an increased risk for CLAD or death (HR 1.52; 95% CI, 0.99–2.34; P = 0.05), with the difference in CLAD-free survival primarily being observed starting at approximately 2.5 years after transplantation.
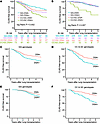 Figure 2
Figure 2Complement-mediated CLAD or death is dependent on DSAs. CLAD-free survival was worse in the recipients with DSAs and the C3 R102G polymorphism (CC/GC). The BJH (A) and UCSF (B) cohorts were stratified by DSA-negative and DSA-positive recipients. The P value was determined by log-rank test. (C and D) Kaplan-Meier plot based on the multivariate Cox regression analysis of CLAD-free survival in the UCSF cohort (C and D) and BJH (E and F), separated by genotype and stratified by DSA status.
These observations also held true in the UCSF cohort, wherein 87 LTx recipients (20.5%) had evidence of DSAs at some point during their post-transplant course. Among participants without DSAs there was no different risk for CLAD or death by the rs2230199 genotype (aHR 1.2; 95% CI, 0.85–1.9; adjusted P = 0.24) (Figure 2B). However, among the participants who developed DSAs and had a CC or a GC allele, there was a 3.2-fold increased risk for CLAD or death (aHR 3.2; 95% CI, 1.6–6.2; adjusted P = 0.0007). These models were adjusted for age, sex, race, transplant diagnosis, and Gram-negative infection. These findings suggest the impact of the rs2230199 SNP is dependent upon DSA development.
We next examined CLAD-free survival separately in recipients based on the presence or absence of C3 R102G. Recipients in the UCSF cohort who lacked C3 R102G (i.e., GG genotype) did not demonstrate a significant difference in CLAD-free survival irrespective of their DSA status (Figure 2C). In comparison, recipients with post-transplant DSAs and the C risk allele (i.e., CG or CC genotype) were at a 3-fold higher risk for worse CLAD-free survival after adjusting for age, sex, race, transplant diagnosis, and Gram-negative infection (Figure 2D). The results also held true in the BJH cohort, in which recipients with post-transplant DSAs and the C risk allele had worse CLAD-free survival (aHR 1.9; 95% CI, 1.1–3.3; adjusted P = 0.02; Figure 2, E and F). These data suggest the worst long-term outcomes occurred in those who developed post-transplant DSAs and harbored the rs2230199 SNP.
Impaired complement regulation promotes CLAD in the mouse orthotopic LTx model. The C3 R102G polymorphism lacks a synonymous variant in mice. However, mice deficient in Crry (Crry–/–) have increased complement activation due to impaired Factor I–mediated cleavage of C3b and C4b, resulting in dysregulated C3 convertase formation on the membrane (38). As a result, these mice have more C3 consumption and lower circulating levels of C3 (39, 42), as well as attenuated levels of C3 in their bronchoalveolar lavage (BAL) (Figure 3A). Given our prior findings that C3 is required for epithelial cell survival (27) and that C3 R102G predisposes to impaired complement regulation (36), we asked if CLAD development is regulated by complement activation. For this purpose, we used a previously established orthotopic mouse LTx model of CLAD (43, 44) (Figure 3B). Wild-type Crry+/+ and Crry–/– recipients, both on a B6 background (H-2b), were engrafted with major-mismatched left lungs encoding 3 transgenes (3T-FVB; H-2q): a reverse tetracycline activator gene driven by the club cell secretory protein promoter, a Cre recombinase gene under the control of the reverse tetracycline activator, and a lox-P activated diphtheria toxin A gene. These transgenes induce graft-specific club cell injury and depletion in B6 recipients following 2.5 days of doxycycline (DOX) ingestion, leading to severe CLAD development with a combination of obliterative and restrictive fibrotic pathology that is dependent on alloimmune responses to a major histocompatibility class mismatch. In contrast, 30 hours of DOX ingestion only induces minimal club cell depletion and results in markedly less allograft injury.
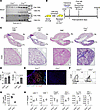 Figure 3
Figure 3Dysregulated complement activation promotes CLAD in a mouse LTx model. (A) Immunoblot of serum and BAL C3-α and -β fragments from resting Crry+/+ and Crry–/– mice (n = 4/group). The data shown are representative results of 2 experiments. (B) Mouse orthotopic left LTx model of CLAD. (C) H&E and Masson’s trichrome (MT) staining of a heart-lung block from POD 16 Crry+/+ and Crry–/– allograft recipients. Scale bars: 250 μm. The histology shown is a representative result of at least 6 transplants per group. (D) POD 16 blinded A (vascular) and B (airway) ISHLT consensus pathological scoring (n = 7/group). (E) immunofluorescent staining of POD 16 allograft club and ciliated cells using Abs specific for CCSP and acetylated tubulin (Ac-Tubulin). The images shown are representative results of 5 transplants per group. (F) A representative flow cytometry plot gating strategy for identifying and (G) quantifying POD 16 CD326+ CCSP+ (club) cells in POD 16 allografts (n = 4/group). (H) POD 16 intragraft total and indicated subset CD4+ and CD8+ T cell numbers (n = 4/group). Bar graphs and dot plots show mean ± SD for (D) Mann-Whitney U test and (G and H) Welch’s t test. *P < 0.05, ***P < 0.001. NL, native lung.
After induction of immunosuppression-mediated lung allograft acceptance, recipients ingested DOX on postoperative day (POD) 7 for 30 hours; on POD 16, allografts were examined for histological evidence of CLAD. We observed more evidence of peribronchiolar fibrosis, obliterative airway disease, scarring of the alveolar parenchyma, and collagen deposition in allografts of Crry–/– when compared with wild-type recipients (Figure 3C and Supplemental Figure 1). Blinded grading for LTx rejection using consensus International Society for Heart and Lung Transplantation (ISHLT) criteria (45) showed significantly higher “B” airway inflammation in allografts after transplantation into Crry–/– relative to wild-type recipients (Figure 3D). Consistent with these findings, club cells failed to fully reconstitute the graft epithelium in Crry–/– hosts. Some club cells were observed scattered throughout the interstitium, suggesting epithelial repair is inhibited by augmented complement activation (Figure 3, E–G). Similar to previous findings in humans with CLAD (46, 47), there were more intragraft Th17 and IFN-γ+ CD8+ T cells in mouse recipients lacking Crry (Figure 3H).
Impaired complement regulation promotes CLAD in a B cell–dependent manner. Previous work has shown that activated C3 fragments bind to B cells to enhance their activation (48, 49). Given these observations, we asked whether Crry deficiency was associated with increased B cell activity in our CLAD model. C3d deposition on B cells was increased in the spleen and allografts of Crry–/– recipients (Figure 4A). There were also more total and proliferating CD19+B220+ B cells in the allografts of Crry–/– recipients (Figure 4B). We also quantified B cell levels in human LTx recipients carrying the C3 R102G variant (Figure 4C). A subcohort of 218 recipients with C3 R102G had increased CD19+ B cell frequencies in their BAL fluid after adjusting for age, sex, diagnostic group, and ethnicity. Likewise, we observed higher numbers of B cells in the BAL fluid of Crry–/– compared with wild-type recipients (Figure 4D and Supplemental Figure 2). De-novo DSAs are associated with an increased risk for CLAD development and progression (50). Therefore, we next measured DSA titers in Crry–/– recipients. Relative to wild-type recipients, Crry–/– recipients had higher IgM, IgG2c, and IgG3 DSAs in their BAL fluid (Figure 4E). By contrast, only IgM DSA numbers were elevated in the peripheral circulation of Crry–/– recipients (Figure 4F). Because DSA accumulation in BAL fluid indicated the local production of Abs, we assessed the abundance of Ab-secreting cells within allograft tissue. Analysis of allografts of Crry–/– recipients revealed more intragraft IgM+ and IgG+ Ab-secreting cells relative to wild-type recipients (Figure 4G and Supplemental Figure 3). CD73, CD86, and CD273 co-expression defines a memory B cell subset poised to become Ab-secreting cells (51). Strikingly, only Crry–/– recipient allografts had IgM+ and IgG+ CD73+CD80+CD273+ memory phenotype B cells (Figure 4, H and I).
 Figure 4
Figure 4Enhanced intragraft B cell accumulation and activation in lung recipients with a defect in complement regulation. (A) Levels of C3d bound to B cells 2 days after the induction of DOX-induced allograft club cell injury. Representative FACS histogram of C3d B cell staining and (right panel) plots of C3d MFI normalized to a corresponding fluorescence minus one (FMO) control (n = 4/group). (B) POD 16 FACS contour plot of intragraft B cell abundance (left), ki67 histogram (proliferation) (middle), and a plot of total allograft B cell numbers on POD 16 (right). FACS contour plots and histograms are representative results from 4/group. (C) Percentages of BAL CD19+ lymphocytes in UCSF cohorts who carry the GG genotype and C3 R102G (CC&GC) polymorphisms. P = 0.02 by an unpaired t test. (D) Quantitation of BAL B cell numbers in POD 16 allografts (n = 4/group). Levels of POD 16 of (E) BAL (n = 4/group) and (F) serum (n = 5/group) donor reactive IgM, IgG1, IgG2c, and IgG3. (G) POD 16, intragraft CD138+ IgM+ and IgG+ Ab-secreting cell numbers (n = 5/group). (H) A representative FACS gating strategy used to identify intragraft CD80+CD73+CD273+ memory B cells, as shown in the bar graphs (I), which display the total intragraft numbers of these cells expressing either surface IgM (n = 5/group) or IgG (n = 4/group) on POD 16. Bar graphs and dot blots show mean ± SD values according to 2-way ANOVA with Šidák’s multiple comparisons test (A), and Welch’s t test (B, D–G, and I). *P < 0.05, **P < 0.01, ***P < 0.001.
Given that our data indicated dysregulated complement regulation triggers the production of DSAs, we next sought evidence of intragraft classical pathway activation by assessing C3d deposition. Relative to wild-type lung recipients, C3d deposition was highly prevalent on B cells within tertiary lymphoid aggregates and, to a lesser degree, on the bronchial epithelium and vascular endothelium in allografts of Crry–/– recipients (Figure 5, A and B). We also analyzed Ab deposition in the lung parenchyma. IgG deposition was more apparent on the allograft alveolar capillaries and bronchial epithelium of Crry–/– recipients relative to the native lungs of Crry–/– recipients or wild-type recipient allografts (Figure 6, A and B). Although IgM expression was present on cells within the pleural cavity, IgM deposition was almost undetectable on lung parenchyma (Supplemental Figure 4).
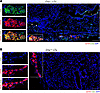 Figure 5
Figure 5Lung allografts from Crry-deficient recipients show elevated C3d deposition. C3d and CD19 immunofluorescent staining of Crry–/– (A) and wild-type Crry+/+ (B) recipient allograft tissue (LTx). Stars and arrows denote C3d staining on vascular endothelium and bronchial epithelium, respectively. Rectangular insets depict tertiary lymphoid aggregates enriched with B cells. Scale bars: 150 μm. The images shown are representative results of at least 3 transplants per group.
 Figure 6
Figure 6IgG deposition is more pronounced on lung allograft parenchyma from Crry-deficient recipients. (A) Immunofluorescence detection of IgG and IgM deposition on alveolar capillaries, as denoted by CD31 Ab staining. Inset scale bars: 25 μm. White arrowheads denote IgG deposition on alveolar capillaries. Images shown are representative of 4 transplants per group. (B) IgG and IgM deposition analysis on allograft bronchioles (b) and vessels (v). Inset scale bars: 150 μm. Images shown are representative of 4 transplants per group.
Finally, to examine if B cells are required for CLAD development in our model, Crry–/– recipients were treated with B cell–depleting Abs and evaluated on POD 16. Compared with recipients that received control Abs, B cell depletion significantly inhibited pathological signs of CLAD development (Figure 7, A and B, and Supplemental Figure 5).
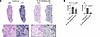 Figure 7
Figure 7CLAD development in Crry-deficient allograft recipients is driven by B cells. (A) Crry-deficient lung recipients received B cell–depleting Abs or control Ig on PODs 6 and 12 and were evaluated for allograft injury on POD 16. Inset scale bars: 200 μm. The histology shown is a representative result of H&E and Masson’s trichrome (MT) allograft staining of 5 transplants per group. (B) POD 16 blinded A (vascular) and B (airway) ISHLT consensus pathological scoring (n = 5/group). The bar graph shows mean ± SD; the Mann-Whitney U test was used to generate P values.
-
Discussion
CLAD is characterized by a persistent and irreversible decline in lung function and is the major cause of long-term morbidity and mortality after LTx (2, 52). Both innate and adaptive immune responses contribute to the progression of CLAD (53). The complement system is an early component of the innate immune response and has been shown to influence outcomes in primary graft dysfunction, a form of acute lung injury occurring early after LTx (54–56). However, complement activation also influences long-term adaptive immune responses in multiple organ systems and has been implicated in CLAD pathogenesis (9). Complement activation has been shown to augment alloimmune CD4+ and CD8+ T cell responses through C3a-C3aR interactions (21, 57, 58). More specifically, IL-17 activation in lung allografts suppresses membrane regulatory protein expression in airway epithelial cells and results in increased local complement activation (9). Increased C3a levels enhances IL-17 production, setting up a feed-forward loop that worsens obliterative bronchiolitis. Increased C3a levels also upregulate TGF-β, a key mediator of obliterative bronchiolitis, which downregulates membrane regulatory proteins such as CD46/Crry and CD55 in models of lung fibrosis (23, 59). This TGF-β–mediated loss of regulatory proteins is abrogated by both C3aR and C5aR1 inhibition, demonstrating how complement activation is a key component of amplification loops propagating irreversible airway and parenchymal fibrosis (23, 59).
There exists considerable variability in the onset of CLAD. Risk factors such as infection, air pollution, aspiration, and a prior history of primary graft dysfunction or acute cellular rejection (ACR) predispose to an earlier onset of CLAD (1, 6, 60). Given the heterogeneity of immune responses, it has been increasingly recognized that particular recipients may also be genetically predisposed to worse CLAD-free survival (61, 62). The C3 R102G functional polymorphism has also been associated with worse outcomes in kidney and liver transplantation (63, 64). LTx recipients who had a donor with C3 R102G had worse bronchiolitis obliterans syndrome–free survival (65). However, at least in renal transplantation, there has been no benefit to matching donors with recipients, based on this polymorphism (66), and long-term LTx outcomes in recipients with advanced lung disease and C3 R102G have not been investigated to date. Surprisingly, we observed that nearly one-third of LTx recipients harbored C3 R102G polymorphism, which exceeds the minor allele frequency of 10% to 15% in the general population, suggesting it may also play a role in the pathophysiology of the underlying lung disease. However, having such a common polymorphism among recipients also necessitates a better understanding of how it affects CLAD-free survival in the setting of impaired complement regulation. Moreover, the prevalence of this functional polymorphism affords avenues to test whether modulating complement activation in LTx recipients with this polymorphism may facilitate personalized immunomodulation to improve CLAD-free survival.
To further probe the effects of dysregulated complement activation on CLAD development, we analyzed graft injury and inflammation in an orthotopic mouse LTx model using Crry–/– mice as recipients. Crry is a membrane complement regulator expressed on a wide variety of murine cells that prevents C3 fragment deposition in response to classical or alternative complement pathway activation (67). Like CD46, Crry regulates complement activation through Factor I–mediated cofactor activity (38, 39). In contrast to wild-type recipients, Crry–/– recipients had considerably more evidence of graft airway injury and higher levels of DSAs, suggesting that CLAD results from Ab-mediated complement activation. However, whether DSA-mediated complement activation is required for graft epithelial injury remains controversial. Introducing either complement-activating or non–complement-activating IgG against donor antigens into a heterotopic mouse tracheal transplant model has been shown to generate obliterative airway disease (68). In the orthotopic mouse LTx model, DSA generation occurred de novo approximately 8 days after club cell injury.
Given that our data indicated dysfunctional complement regulation triggers the production of DSAs, we next sought evidence of intragraft classical pathway activation by assessing C3d deposition. We observed more C3d deposition in Crry–/– compared with wild-type recipient allografts, with most activity concentrated on B cells within tertiary lymphoid aggregates and, to a lesser extent, on the vascular endothelium and bronchial epithelium. In contrast, C3d deposition was nearly absent from allografts of wild-type recipients. Additionally, IgG deposition was more apparent on the allograft alveolar capillaries and bronchial epithelium in the setting of impaired complement regulation in recipients. Although IgM expression was present on cells within the pleural cavity, IgM deposition was almost undetectable on lung parenchyma. Finally, we showed that B cell depletion significantly inhibited pathological signs of CLAD development in our animal model. Hence, despite the intrinsic differences in the animal model of Crry deficiency compared with the C3 polymorphism in humans, the work aids us in deciphering how impaired complement regulation drives chronic rejection after LTx. Specifically, the C3 polymorphism in humans results in impaired complement regulation as the charge of the C3 protein synthesized in the setting of the R102G polymorphism impairs the ability of the regulatory protein Factor H to inactivate it (36). Recipients harboring this polymorphism had worse CLAD-free survival in the presence of DSAs and had increased numbers of CD19+ B cells in their BAL. In the mouse model, Crry deficiency also resulted in impaired regulation and increased C3 activation (67), thereby leading to accelerated CLAD via a B cell–dependent process. Taken together, these data point to potential pathways by which impaired complement regulation enhances humoral immunity in the setting of LTx.
Increased complement activation has been reported to enhance B cell responses via multiple pathways (69). For example, resting B cells express high amounts of the complement receptor type 2 (CR2/CD21), which binds antigen-bound C3 fragments (48, 70). Co-clustering CR2 with the B cell receptor lowers the threshold of B cell activation, leading to enhanced Ab production (71). CR2 is also expressed on stromal cells and follicular dendritic cells, where it plays a vital role in antigen retention to promote the generation of B cell memory and Ab-secreting cells (72). In this regard, we noted C3d deposition on B cells within tertiary lymphoid aggregates within Crry–/– allografts. Our previous work has demonstrated that germinal-center B cells primarily housed within intragraft tertiary lymphoid tissues play a crucial role in the local production of IgG, but not IgM DSAs, while also promoting the generation of memory B cells and Ab-secreting cells within allograft tissue (73). This raises the possibility that our observation of BAL DSA accumulation in Crry–/– recipients is due to complement-mediated activation of germinal-center B cells (74). To this end, previous work has shown that complement fragment deposition on B cells promotes their activation (75). The B cell receptor C1q binding domain drives the fixation of C3 fragments to the B cell membrane and generates optimal antigen-specific responses (49). Relative to these previous reports, we observed increased B cell accumulation, enhanced proliferative responses, and elevated levels of memory B cells and Ab-secreting cells in allografts after transplantation into Crry–/– recipients. Interestingly, a recent single-cell RNA-sequencing study revealed an association between intragraft accumulation of plasma cells and CLAD in human and mouse allografts (15). Collectively, data from the orthotopic mouse LTx model implicate dysregulated complement activation in stimulating local B cell activation responses that promote CLAD development.
Observations from the animal model prompt questions as to whether the increased susceptibility to CLAD in the setting of the C3 polymorphism is due to B cell–dependent pathways. We observed an increased proportion of BAL CD19+ B cells in recipients with the polymorphism. Moreover, CLAD-free survival in the setting of C3 R102G polymorphism occurred primarily in patients who developed anti-HLA donor-specific Abs, which are B cell dependent. Hence, although DSA prevalence did not differ among those with and without the polymorphism, we attribute these observations to inherent differences in investigating Ab-mediated alloimmune responses between mice and humans. Our categorization of DSAs in patients is based on MFI from a bead-based platform, whereas we measure titers in the mouse model. Other possibilities include contributions from non-HLA Abs (12) and DSAs deposited in the allograft (76, 77). Knowing that RAS is a form of CLAD associated with humoral immune responses and worse outcomes, we also interrogated if patients with the C3 polymorphism had an increased odds of RAS using forced vital capacity as a prognostic adjunct (78), but we did not observe a difference. Moreover, because the B cell enumeration in our patient cohort was done using a clinical flow cytometry panel (79) and we do not have banked BAL samples for this cohort, we were unable to conclusively link the increased BAL CD19+ B cells in our recipients with the polymorphism to B cell activation. Yet, plasma from patients harboring this polymorphism has been shown to activate the alternative pathway (AP) more efficiently, and these patients demonstrate increased AP amplification (36). This increased activation and amplification would generate more complement activation fragments (e.g., C3d), which would bind to receptors such as CR2, thus enhancing B cell responses, including DSA production. Anti-HLA Abs have also been shown to induce epithelial cell proliferation as well as induce the production of soluble growth factors, thus perpetuating injury that leads to CLAD (80, 81). Thus, the worse CLAD-free survival in patients with post-transplant DSAs harboring the C3 R102G polymorphism, and the enrichment of BAL CD19+ B cells, encourage us to build on our observations from the animal model of Crry deficiency in recipients that demonstrated high BAL C3d+ B cells, elevated levels of intragraft DSAs, and B cell–dependent CLAD development. Our data collectively suggest that CLAD development in the setting of dysregulated complement activation is mediated by increased humoral responses. Future work would involve measuring markers of complement activation in the BAL of recipients harboring this polymorphism, improved subphenotyping of B cells and their subsets to assess for complement-mediated activation in longitudinal samples from LTx recipients, as well as staining lung tissues with C3d and IgG, as we have demonstrated in our animal model, to conclusively prove that the C3 R102G polymorphism increases the risk of CLAD via B cell activation.
There are also other potential mechanisms by which complement activation may be increasing CLAD risk, including complement activation–driven T cell effector responses that could indirectly influence B cell responses, increased Ab-dependent cellular cytotoxicity, and direct toxicity through HLA ligation that drives fibrosis (82, 83). From the standpoint of leveraging this polymorphism for patient selection in future clinical trials, mechanistic studies could involve determining whether the C3 R102G mutation worsens allograft injury via a C3d-CR2–mediated axis, perhaps by promoting memory B cell responses. Further investigation into probing impaired classical pathway activation on allograft endothelial surfaces could be addressed with inhibitors of C1-esterase, C3, or C5. Given the availability of FDA-approved anticomplement agents that target these molecules, such therapies could be considered for complement-mediated injury in LTx recipients (16).
Our study has limitations, including insufficient sample sizes at both sites to test whether this polymorphism predisposes to AMR as a mediator of CLAD. Additionally, we did not observe significant differences in the prevalence of ACR or lymphocytic bronchiolitis in either the BJH or the UCSF cohort, regardless of the polymorphism. However, we are cautious about overinterpreting ACR data in human recipients, because pathologist agreement is low, protocols differ between centers, and ACR is a heterogeneous process prone to sampling error (84, 85). Moreover, many patients with ACR do not develop CLAD, and many recipients with CLAD have never experienced prior ACR (11, 86, 87). Emerging data suggest ACR and AMR may coexist within the same allograft and be mechanistically linked (88, 89). Because we cannot exclude the possibility that the C3 polymorphism drives ACR and lymphocytic bronchiolitis, we elected not to include them in the final multivariable models, to prevent collider bias. We also do not routinely measure other non-HLA anti-allograft Abs in the allograft or circulation. However, we observed that IgG2c and IgG3 (but not IgM or IgG1) levels were elevated in the BAL of Crry–/– recipients as compared with in their serum, whereas only IgM was elevated in the serum of Crry–/– recipients compared with wild-type allograft recipients. Along these lines, although IgG1 strongly activates complement in humans, IgG2 subclasses and IgG3 strongly activate complement in mice (90–92). These observations support the role of local Ab production and intragraft complement-mediated responses in LTx recipients as downstream responses to impaired complement regulation. Nevertheless, despite observing the propensity toward CLAD in both humans and mice, we acknowledge our modeling of impaired complement regulation in mice is not entirely analogous to mechanisms that operate in humans (93). Another limitation is that collagenase digestion was used to generate allograft single-cell suspensions, which could cause CD138 shedding, leading to an underestimate of Ab-secreting cell numbers (94). Although we did not include isograft data in our experiments, we have previously reported that syngeneic transplants, in which 3T-B6 lungs were engrafted into B6 recipients, did not develop chronic rejection despite extensive club cell injury (44). Additionally, because there is no validated scoring system for the pathology of CLAD in mouse models, we used the B score for histopathological scoring of airway involvement (95). Moreover, we used Masson’s trichrome staining and tissue hydroxyproline assays as a surrogate for the fibrosis that occurs in chronic rejection. The proportion of recipients with RAS was low to draw any meaningful conclusions, and we did not have total lung capacity data on our recipients in these cohorts. However, whereas increased humoral responses are associated with RAS, DSAs remain a risk for obstructive CLAD (96–98) and can affect lung function even in the absence of clinical AMR (99). Future clinical studies should focus on determining if complement-mediated humoral immune responses contribute to RAS (14).
Using an orthotopic mouse model of LTx, we show that dysregulated complement regulation promotes CLAD through enhancing B cell activation. In a multicenter study, we demonstrate worse CLAD-free survival in recipients who carry the C3 R102G polymorphism and have DSA. The mechanistic basis for this finding may primarily be B cell dependent, suggesting that conventional T cell immunosuppression regimens may be of limited efficacy at preventing CLAD in patients with the C3 R102G polymorphism. By describing a sizeable subgroup of LTx recipients with a polymorphism that predisposes them to impaired complement regulation, we identify a cohort that can form the basis of clinical trials in personalized immunomodulation to improve long-term outcomes after LTx.
-
Methods
Sex as a biological variable. Our study investigated data from both male and female human participants, and similar findings are reported for both sexes. Our orthotopic LTx model used male donor lungs transplanted into male recipients or female donor lungs transplanted into female recipients.
Study design, settings, and participants. We performed a retrospective cohort study and screened 1,218 primary LTx recipients at Washington University/BJH between January 1, 2005, and December 31, 2022, with follow-up through December 31, 2023. We excluded those who did not consent (n = 250) and those for whom genotyping had not been performed (n = 576) (Figure 1A). We also included 864 primary LTx recipients at UCSF through September 30, 2021. We excluded those who did not consent (n = 153) and those for whom genotyping had not been performed (n = 293) (Figure 1B).
Clinical variables. Clinical management at these 2 centers has been described in prior studies (11, 61). HLA Abs were detected using the LABScreen Single Antigen assay, and DSA testing was considered positive if the reported MFI was ≥2,000. The time from transplantation to the first detection of DSA after transplantation was defined as time to DSA positivity. A positive bacterial culture from a bronchial wash or BAL was defined as bacterial isolation. Primary graft dysfunction and ACR were defined based on ISHLT criteria. CLAD was defined as a persistent decline in forced expiratory volume in 1 second ≤80% for at least 3 weeks without a specific cause. CLAD-free survival was defined as the earliest occurrence of either CLAD or death, whichever came first.
Genotyping. At BJH, salivary specimens from consented patients were genotyped for the C3 R102G (rs2230199) polymorphism, based on the TaqMan assay, as previously described (100). At UCSF, blood specimens were genotyped using a transplant-targeted Affymetrix gene array designed by the iGeneTRAiN consortium that directly typed the C3 R102G (rs2230199) polymorphism (101). At both centers, recipients with CC/CG (minor allele) were compared with those with GG.
BAL analysis. Specimens included in this study were from bronchoscopies conducted as part of routine clinical management, when patients had an unexplained decline in their lung function and were being evaluated for CLAD. In the UCSF cohort, bronchoscopies were performed for clinical indication or for allograft surveillance at 0.5, 1, 2, 3, 6, 12, 18, and 24 months after transplantation. BAL cell subsets, including B cells, are routinely quantified by clinical cytometry at UCSF.
Crry–/– mice. Crry deficiency has previously been reported as being embryonic lethal because breeding Crry+/– mice together failed to achieve any Crry–/– mice (102–104). However, strategic breeding procedures led to the generation of Crry–/– mice. In detail, breeding Crry+/–C3–/– mice resulted in Crry–/–C3–/– offspring, indicating that Crry–/– mice can survive in the condition of C3 deficiency. We then bred Crry+/– males with Crry–/–C3–/– females to generate Crry–/–C3+/– mice. These mice only have half capacity of the AP activity (39), so we predicted that mothers with the genotype of Crry–/–C3+/– would have insufficient AP activity to attack embryos that had a Crry–/– genotype. Considering this, we bred Crry+/– males with Crry–/–C3+/– females, leading to the generation of Crry–/–C3+/+ pups, which are Crry–/– but C3 sufficient. BAL was performed on these mice by instilling 1 mL of PBS plus enzyme inhibitor (Halt Protease and Phosphatase Inhibitor Single-Use Cocktail; Thermo Fisher, catalog 78442) into both lungs, performing centrifugation at 500g for 5 minutes, followed by aliquoting the supernatant.
Immunoblot analysis. C3 was detected using immunoblotting of BAL and serum (1:3,000; MP Biomedicals, catalog 55444). BAL supernatant (3 μL) was mixed with 7 μL of PBS, resulting in a total volume of 10 μL. Subsequently, 3.5 μL of 4× reducing buffer (1:10 ratio of β-mercaptoethanol added to Laemmli buffer; Bio-Rad, catalog 1610747) was added to the diluted samples, bringing the final volume to 13.5 μL. The samples were vortexed for 5 seconds and then centrifuged. After centrifugation, the samples were incubated in a heating block at 98°C for 6 minutes. After this heating step, the samples were centrifuged again before being loaded onto the gel.
For serum samples, 1 μL of each sample was combined with 9 μL of PBS, also resulting in a total volume of 10 μL. Like the BAL samples, 3.5 μL of 4× reducing buffer was added, and the samples were vortexed, centrifuged, heated, and centrifuged again before gel loading. Purified C3 protein was prepared at a final concentration of 7.5 ng/μL by diluting 1 μL of the purified protein in 99 μL of PBS, followed by the addition of 33.5 μL of 4× reducing buffer.
Orthotopic left LTx. C57BL/6 (B6) mice were purchased from The Jackson Laboratories. Club cell secretory protein (CCSP) rtTA/TetOCre/DT-A mice on a mixed background were initially generated by Jeffrey Whitsett of The Children’s Hospital of Cincinnati. These mice were then backcrossed by our group to greater than 99% onto a FVB/J background using microsatellite-assisted accelerated backcrossing (MAX-BAX; Charles River) to generate 3T-FVB left lung donors (44). Left lung orthotopic LTx has been described (105).
To induce allograft acceptance, recipients received 250 μg of CD40L Abs (clone MR1) i.p. on POD 0 and 200 μg of human recombinant CTLA4 Ig on POD2 (106). Club cell injury was triggered by DOX ingestion via food (625 mg/kg chow; ENVIGO) and water (2 mg/mL; Doxycycline hyclate, MilliporeSigma) for 30 hours to induce club cell injury. To deplete B cells, rat anti–mouse CD19 (clone 1D3; Bio-X-Cell), rat anti–mouse B220 (clone RA3.3A1/6.1; Bio-X-Cell), and mouse anti–mouse CD22 (clone CY34.1; Bio-X-Cell) (150 μg/mouse for each) were administered i.p. on PODs 6 and 12. Control recipients received 450 μg of rat IgG1 isotype control (clone TNP6A7; Bio-X-Cell). After 1 day, recipients were injected with mouse anti–rat IgG (clone Mar-18.5; Bio-X-Cell) at 150 μg/mouse.
Immunohistological staining. Harvested grafts were formaldehyde fixed, paraffin embedded, and stained with H&E or Masson’s trichrome stain. LTx histology was graded by a blinded pathologist using the 2007 revision of the 1996 ISHLT working formulation for the standardization of nomenclature in the diagnosis of lung rejection (45). For immunohistochemical analysis, paraffin sections were first blocked with 5% goat serum and 2% fish gelatin (both from Sigma-Aldrich) at 25°C for 45 minutes. Sections were then stained with 1:500 polyclonal rabbit anti–mouse/rat CCSP (catalog WRAB-3950, Seven Hills Bioreagents) and mouse anti-acetylated tubulin (1:5,000; clone 6-11 B-1; Sigma-Aldrich) overnight at 4°C. For secondary Ab-mediated immunofluorescent visualization, we used 1:1,000 goat anti–mouse Alexa Fluor 488–labeled secondary Abs (catalog A-11-001, Thermo Fisher), 1:1,000 donkey anti–goat Alexa Fluor 488 (catalog A-11055, Thermo Fisher), 1:1,000 donkey anti–rabbit Alexa Fluor 555 (catalog A-31572, Thermo Fisher), and 1:1,000 goat anti–rabbit Alexa Fluor 555 (catalog 4413S, Cell Signaling Technology). To assess C3d tissue deposition, 1:100 goat-anti–mouse/rat C3d (Bio-Techne), 1:100 rabbit anti–goat IgG-biotin (catalog 31172, Thermo Fisher), dilutions were used and stained with VectorStain ABC-HRP kit (catalog PK-4000, Vector) according to the manufacturer’s recommendations. To detect IgM and IgG adsorption to lung parenchyma, allografts were first subjected to three 1.0 mL PBS BAL extractions, flushed vascularly to remove leukocytes and nonspecifically bound immunoglobulin, paraffin embedded, and then stained with 1:250 anti-mouse CD31-FITC (clone 330, Thermo Fisher), 1:100 goat anti–mouse IgG-PE (catalog 31861, Thermo Fisher) or 1:100 goat anti–mouse IgG-FITC (catalog 31457, Thermo Fisher), and 1:100 goat anti–mouse IgM-DyeLight 650 (catalog SA5-10153, Thermo Fisher) or 1:100 goat anti–mouse IgM-Alexa Flour 555 (catalog A-21426, Thermo Fisher).
Hydroxyproline analysis. To determine intragraft collagen concentrations, a Hydroxyproline Assay Kit (catalog MAK569, Sigma-Aldrich) was used according to the manufacturer’s recommendations. Allograft tissue was extracted for protein with 6.0 N HCl for 3 hours at 120°C and dried at 60°C overnight. Then, 10 μg of protein was assayed for reactivity for oxidized hydroxyproline with 4-(dimethylamino) benzaldehyde, which yields a colorimetric product that was quantified by absorbance at 560 nm on a Biotek Synergy HTX Microplate Analyzer.
Flow cytometric analysis. Lung tissue was minced and digested in an RPMI 1640 solution with type 2 collagenase (50 μg/mL; Worthington Biochemical Corp.) and 5 units/mL DNAse (MilliporeSigma) for 30 minutes at 37°C and then filtered through a 70 μm cell strainer (Thermo Fisher) and treated with ammonium-chloride-potassium lysing buffer (Worthington Biochemical Corp.). Live cell discrimination was conducted with Zombie Fixable Dye (BioLegend). Cell surface staining was performed with the following Abs: CD45 (clone 30-F11; eBioscience), CD45.2 (clone 104; BioLegend), CD90.2 (clone 53-2.1; eBioscience), CD4 (clone RM4-5; eBioscience), CD8α (clone 53-6.7; eBioscience), B220 (RA3-6B2; Biolegend), CD19 (1D3; BD Biosciences), rabbit anti–human/mouse/rat C3d (catalog bs-4877R; Bioss), CD138 (281-2; Biolegend) CD38 (S21016F; Biolegend), IgD (11-26c.2a; Biolegend), Foxp3 (FJK-16s; Thermo Fisher), CD73 (TY/11.8; Biolegend), CD80 (16-10A1; Biolegend), CD273 (TY25; Thermo Fisher), CD31 (clone 390; BioLegend), CD34 (clone HM34; BioLegend), and CD326 (clone G8.8; BioLegend). Intracellular CCSP (Seven Hills Bioreagents) and polyclonal goat anti–mouse IgM heavy chain (SouthernBiotech) staining was conducted with the Cytofix/Cytoperm kit (BD Biosciences) following the manufacturer’s recommendations. Staining for Foxp3 (FJK-16s; eBioscience) and Ki-67 (16A8; BioLegend) was conducted with the Intranuclear Transcription Factor Staining Buffer Kit (Invitrogen) per the manufacturer’s recommendations. For IFN-γ and IL-17A expression, cells were first stimulated with 1 μM ionomycin (MilliporeSigma) and 20 ng/mL PMA (MilliporeSigma) for 3.5 hours, with 2 μM Golgi Plug (BD Biosciences) added for the last 3 hours of stimulation. Cells were then intracellularly stained with IFN-γ (clone XMG1.2; eBioscience) and IL-17A (clone TC11-18H10.1; BioLegend) using a Cytofix/Cytoperm kit. For DSA analysis, FVB splenocytes were resuspended at 4 × 106 cells/mL in FACS buffer (PBS, 0.1% BSA, 0.02% NaN3) and treated with FcγR block on ice for 10 minutes (clone 93; Biolegend). BAL fluid was diluted to 1:15 in PBS, and serum was diluted to 1:25 in PBS, and 50 μL of these diluted specimens was mixed with 50 μL of FVB splenocytes, which were then incubated for 1 hour at 4°C. Cells were washed twice with FACS buffer and stained with goat anti–mouse IgM heavy chain-APC (catalog 1020-11S, SouthernBiotech), goat-anti–mouse IgG2c-AlexaFluor 647 (catalog 1079-3, Southern Biotech), goat-anti–mouse IgG1–Alexa Fluor 488 (catalog 115-545-071, Jackson ImmunoResearch), and goat anti–mouse IgG or goat anti–mouse IgG3 Alexa Fluor 488 (catalog A-21151, Thermo Fisher), and then washed twice more before FACS-based MFI analysis.
Statistics. Data were analyzed by the Mann-Whitney U test or 1-way ANOVA and are represented as mean ± SD. Statistical analysis was conducted in R (version 4.3.2) and GraphPad Prism software, version 9.0. P < 0.05 was considered significant. Time-to-event analyses were performed using Kaplan-Meier curves with log-rank test for equality. Multivariable analyses were conducted via Cox proportional hazards regression.
Study approval. Animal experiments were conducted in accordance with an approved IACUC protocol (Washington University, 19-0827). The BJH participants provided written informed consent in accordance with the Washington University School of Medicine Institutional Review Board for Human Studies protocol (no. 201811073 and 201105421). The UCSF participants provided written informed consent in accordance with the Institutional Review Board for Human Studies protocol (no. 13-10738).
Data availability. Values for all data points in graphs are reported in the Supporting Data Values file. The study participants have not consented to sharing genetic data publicly; therefore, the data cannot be uploaded to a publicly accessible server. Deidentified data are provided using a material transfer agreement/data use agreement with the institutions.
-
Author contributions
HSK, DRC, JRG, and AEG designed the studies. HSK, FL, ZL, AT, LG, DHK, XW, and AEG conducted the experiments. HSK, LKT, DRC, FL, ZL, LG, NRS, CC, HJH, CAW, RRH, DZ, YT, and VED acquired the data. HSK, LKT, DRC, CTB, and AEG analyzed the data. XW, DHK, DEB, DK, and JPA provided reagents. HSK, LKT, DRC, LG, XW, JRG, and AEG wrote the manuscript. HSK, LKT, DRC, DK, JPA, JRG, and AEG acquired funding for this work.
-
Funding support
This work is the result of NIH funding, in whole or in part, and is subject to the NIH Public Access Policy. Through acceptance of this federal funding, the NIH has been given a right to make the work publicly available in PubMed Central.
- NIH grants R01-HL166449, R01-HL169860, and R35-GM136352 to HSK.
- Children’s Discovery Institute to HSK.
- Longer Life Foundation to HSK.
- NIH grant K01-HL155231 to LKT.
- Robert Wood Johnson Foundation to LKT.
- Doris Duke Charitable Foundation to LKT.
- VA Office of Research and Development grant BX005301 to DRC.
- Cystic Fibrosis Foundation grant to CALABR22G0 DRC.
- NIH grants P01-AI116501 and R01HL094601 to DK.
- US Department of Veteran’s Affairs grant 1I01BX002730) to DK.
- Cystic Fibrosis Foundation grant KREISE24AB0-CLAD to DK.
- Foundation for Barnes-Jewish Hospital to DK.
- NIH grant R35-GM136352 to JPA and XW.
- NIH grants R01-HL161048, R01-HL151552 to JRG.
- Cystic Fibrosis Foundation grant GREENL21AB0 to JRG.
- VA Office of Research and Development grant CX002011 to JRG.
- NIH grants P01-AI116501, R01HL167277, R01HL094601, R01HL157407, and P41EB025815 to AEG.
- Cystic Fibrosis Foundation grant GELMAN25XX0 to AEG.
- The Barnes-Jewish Foundation to AEG.
-
Acknowledgments
The authors thank Dorit Daphna-Iken, Jihong Zhu, Rafael Aponte Alburquerque, and Ayse Naz Ozanturk for their technical assistance.
Address correspondence to: Hrishikesh Kulkarni, Division of Pulmonary, Critical Care and Sleep Medicine, University of California, Los Angeles, 10833 Le Conte Ave, 52-257 CHS, Mail code: 169017, Los Angeles, California 90095, USA. Email: hskulkarni@mednet.ucla.edu. Or to: Andrew E. Gelman, Mary Culver Department of Surgery, Division of Cardiothoracic Surgery, Washington University School of Medicine, 660 S Euclid Ave, St. Louis, Missouri 63110, USA. Phone: 314.362.8382; Email: agelman@wustl.edu.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Copyright: © 2025, Kulkarni et al. This is an open access article published under the terms of the Creative Commons Attribution 4.0 International License.
Reference information: J Clin Invest. 2026;136(1):e188891.https://doi.org/10.1172/JCI188891.
-
References
- DerHovanessian A, et al. Chronic lung allograft dysfunction: evolving concepts and therapies. Semin Respir Crit Care Med. 2018;39(2):155–171.
- Venado A, et al. Chronic lung allograft dysfunction. Thorac Surg Clin. 2022;32(2):231–242.
- Kulkarni HS, et al. Bronchiolitis-obliterans syndrome-free survival of lung transplant recipients — analysis of the ISHLT registry. J Heart Lung Transplant. 2017;36(4):310–311.
- Verleden GM, et al. Chronic lung allograft dysfunction: Definition, diagnostic criteria, and approaches to treatment-A consensus report from the Pulmonary Council of the ISHLT. J Heart Lung Transplant. 2019;38(5):493–503.
- Vandermeulen E, et al. Humoral immunity in phenotypes of chronic lung allograft dysfunction: A broncho-alveolar lavage fluid analysis. Transpl Immunol. 2016;38:27–32.
- Sato M. Bronchiolitis obliterans syndrome and restrictive allograft syndrome after lung transplantation: why are there two distinct forms of chronic lung allograft dysfunction? Ann Transl Med. 2020;8(6):418.
- Ali HA, et al. Complement system in LT. Clin Transplant. 2018;32(4):e13208.
- Kallio EA, et al. Blockade of complement inhibits obliterative bronchiolitis in rat tracheal allografts. Am J Respir Crit Care Med. 2000;161(4 pt 1):1332–1339.
- Suzuki H, et al. Role of complement activation in obliterative bronchiolitis post-lung transplantation. J Immunol. 2013;191(8):4431–4439.
- Hachem RR, et al. Human leukocyte antigens antibodies after lung transplantation: Primary results of the HALT study. Am J Transplant. 2018;18(9):2285–2294.
- Kulkarni HS, et al. Pseudomonas aeruginosa and acute rejection independently increase the risk of donor-specific antibodies after lung transplantation. Am J Transplant. 2020;20(4):1028–1038.
- Xu Q, et al. Chronic lung allograft dysfunction is associated with an increased number of non-HLA antibodies. J Heart Lung Transplant. 2024;43(4):663–672.
- Watanabe T, et al. A B cell-dependent pathway drives chronic lung allograft rejection after ischemia-reperfusion injury in mice. Am J Transplant. 2019;19(12):3377–3389.
- Misumi K, et al. Humoral immune responses mediate the development of a restrictive phenotype of chronic lung allograft dysfunction. JCI Insight. 2020;5(23):e136533.
- Smirnova NF, et al. Single-cell transcriptome mapping identifies a local, innate B cell population driving chronic rejection after lung transplantation. JCI Insight. 2022;7(18):e156648.
- Kulkarni DH, et al. Local complement activation and modulation in mucosal immunity. Mucosal Immunol. 2024;17(4):739–751.
- Babu AN, et al. Microvascular destruction identifies murine allografts that cannot be rescued from airway fibrosis. J Clin Invest. 2007;117(12):3774–3785.
- Hodge S, et al. Decreased efferocytosis and mannose binding lectin in the airway in bronchiolitis obliterans syndrome. J Heart Lung Transplant. 2011;30(5):589–595.
- Magro CM, et al. Association of humoral immunity and bronchiolitis obliterans syndrome. Am J Transplant. 2003;3(9):1155–1166.
- Schaapherder AF, et al. Porcine islet cells of Langerhans are destroyed by human complement and not by antibody-dependent cell-mediated mechanisms. Transplantation. 1996;62(1):29–33.
- Vieyra M, et al. Complement regulates CD4 T-cell help to CD8 T cells required for murine allograft rejection. Am J Pathol. 2011;179(2):766–774.
- Gu H, et al. Contribution of the anaphylatoxin receptors, C3aR and C5aR, to the pathogenesis of pulmonary fibrosis. FASEB J. 2016;30(6):2336–2350.
- Gu H, et al. Crosstalk between TGF-β1 and complement activation augments epithelial injury in pulmonary fibrosis. FASEB J. 2014;28(10):4223–4234.
- Liszewski MK, et al. Intracellular complement activation sustains T cell homeostasis and mediates effector differentiation. Immunity. 2013;39(6):1143–1157.
- King BC, et al. Complement component C3 is highly expressed in human pancreatic islets and prevents β cell death via ATG16L1 interaction and autophagy regulation. Cell Metab. 2019;29(1):202–210.
- Kulkarni HS, et al. Intracellular C3 protects human airway epithelial cells from stress-associated cell death. Am J Respir Cell Mol Biol. 2019;60(2):144–157.
- Sahu SK, et al. Lung epithelial cell-derived C3 protects against pneumonia-induced lung injury. Sci Immunol. 2023;8(80):eabp9547.
- Elvington M, et al. A C3(H20) recycling pathway is a component of the intracellular complement system. J Clin Invest. 2017;127(3):970–981.
- Forneris F, et al. Structures of C3b in complex with factors B and D give insight into complement convertase formation. Science. 2010;330(6012):1816–1820.
- Budding K, et al. A promoter polymorphism in the CD59 complement regulatory protein gene in donor lungs correlates with a higher risk for chronic rejection after lung transplantation. Am J Transplant. 2016;16(3):987–998.
- Liszewski MK, et al. Complement dysregulation and disease: insights from contemporary genetics. Annu Rev Pathol. 2017;12:25–52.
- Matsuo S, et al. In vivo effects of monoclonal antibodies that functionally inhibit complement regulatory proteins in rats. J Exp Med. 1994;180(5):1619–1627.
- Nagano F, et al. Expression of a Crry/p65 is reduced in acute lung injury induced by extracellular histones. FEBS Open Bio. 2022;12(1):192–202.
- Atkinson C, et al. Targeted complement inhibition by C3d recognition ameliorates tissue injury without apparent increase in susceptibility to infection. J Clin Invest. 2005;115(9):2444–2453.
- Li C, et al. A novel injury site-natural antibody targeted complement inhibitor protects against lung transplant injury. Am J Transplant. 2021;21(6):2067–2078.
- Heurich M, et al. Common polymorphisms in C3, factor B, and factor H collaborate to determine systemic complement activity and disease risk. Proc Natl Acad Sci U S A. 2011;108(21):8761–8766.
- Spencer KL, et al. C3 R102G polymorphism increases risk of age-related macular degeneration. Hum Mol Genet. 2008;17(12):1821–1824.
- Kim YU, et al. Mouse complement regulatory protein Crry/p65 uses the specific mechanisms of both human decay-accelerating factor and membrane cofactor protein. J Exp Med. 1995;181(1):151–159.
- Wu X, et al. Membrane protein Crry maintains homeostasis of the complement system. J Immunol. 2008;181(4):2732–2740.
- Karczewski KJ, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581(7809):434–443.
- Snyder LD, et al. Implications for human leukocyte antigen antibodies after lung transplantation: a 10-year experience in 441 patients. Chest. 2013;144(1):226–233.
- Ruseva MM, et al. Crry deficiency in complement sufficient mice: C3 consumption occurs without associated renal injury. Mol Immunol. 2009;46(5):803–811.
- Liu Z, et al. Reprogramming alveolar macrophage responses to TGF-β reveals CCR2+ monocyte activity that promotes bronchiolitis obliterans syndrome. J Clin Invest. 2022;132(19):e159229.
- Liu Z, et al. An obligatory role for club cells in preventing obliterative bronchiolitis in lung transplants. JCI Insight. 2019;5(9):e124732.
- Stewart S, et al. Revision of the 1996 working formulation for the standardization of nomenclature in the diagnosis of lung rejection. J Heart Lung Transplant. 2007;26(12):1229–1242.
- Burlingham WJ, et al. IL-17-dependent cellular immunity to collagen type V predisposes to obliterative bronchiolitis in human lung transplants. J Clin Invest. 2007;117(11):3498–3506.
- Hodge G, et al. BOS is associated with increased cytotoxic proinflammatory CD8 T, NKT-Like, and NK cells in the small airways. Transplantation. 2017;101(10):2469–2476.
- Qin D, et al. Evidence for an important interaction between a complement-derived CD21 ligand on follicular dendritic cells and CD21 on B cells in the initiation of IgG responses. J Immunol. 1998;161(9):4549–4554.
- Rossbacher J, Shlomchik MJ. The B cell receptor itself can activate complement to provide the complement receptor 1/2 ligand required to enhance B cell immune responses in vivo. J Exp Med. 2003;198(4):591–602.
- Morrell MR, et al. De novo donor-specific HLA antibodies are associated with early and high-grade bronchiolitis obliterans syndrome and death after lung transplantation. J Heart Lung Transplant. 2014;33(12):1288–1294.
- Tomayko MM, et al. Cutting edge: Hierarchy of maturity of murine memory B cell subsets. J Immunol. 2010;185(12):7146–7150.
- Christie JD, et al. Lung transplantation. N Engl J Med. 2024;391(19):1822–1836.
- Jeong JC, et al. Update on the immunological mechanisms of primary graft dysfunction and chronic lung allograft dysfunction. Curr Opin Organ Transplant. 2024;29(6):412–419.
- Kulkarni HS, et al. Local complement activation is associated with primary graft dysfunction after lung transplantation. JCI Insight. 2020;5(17):e138358.
- Shah RJ, et al. Plasma complement levels are associated with primary graft dysfunction and mortality after lung transplantation. Am J Respir Crit Care Med. 2014;189(12):1564–1567.
- Yang W, et al. IL-1β-dependent extravasation of preexisting lung-restricted autoantibodies during lung transplantation activates complement and mediates primary graft dysfunction. J Clin Invest. 2022;132(20):e157975.
- Mathern DR, et al. Absence of recipient C3aR1 signaling limits expansion and differentiation of alloreactive CD8+ T cell immunity and prolongs murine cardiac allograft survival. Am J Transplant. 2019;19(6):1682–1640.
- Werfel T, et al. Activated human T lymphocytes express a functional C3a receptor. J Immunol. 2000;165(11):6599–6605.
- Vittal R, et al. Overexpression of decay accelerating factor mitigates fibrotic responses to lung injury. Am J Respir Cell Mol Biol. 2022;67(4):459–470.
- Amubieya O, et al. Ambient air pollution exposure and outcomes in patients receiving lung transplant. JAMA Netw Open. 2024;7(10):e2437148.
- Calabrese DR, et al. Dectin-1 genetic deficiency predicts chronic lung allograft dysfunction and death. JCI Insight. 2019;4(22):e133083.
- Palmer SM, et al. Innate immunity influences long-term outcomes after human lung transplant. Am J Respir Crit Care Med. 2005;171(7):780–785.
- Brown KM, et al. Influence of donor C3 allotype on late renal-transplantation outcome. N Engl J Med. 2006;354(19):2014–2023.
- Valero-Hervás DM, et al. Complement C3F allotype synthesized by liver recipient modifies transplantation outcome independently from donor hepatic C3. Clin Transplant. 2017;31(1):e12866.
- Kardol-Hoefnagel T, et al. A single nucleotide C3 polymorphism associates with clinical outcome after lung transplantation. Front Immunol. 2019;10:2245.
- Varagunam M, et al. C3 polymorphisms and allograft outcome in renal transplantation. N Engl J Med. 2009;360(9):874–880.
- Foley S, et al. Mouse Crry/p65 is a regulator of the alternative pathway of complement activation. Eur J Immunol. 1993;23(6):1381–1384.
- Takenaka M, et al. Complement activation is not required for obliterative airway disease induced by antibodies to major histocompatibility complex class I: Implications for chronic lung rejection. J Heart Lung Transplant. 2012;31(11):1214–1222.
- Dunkelberger JR, Song W-C. Complement and its role in innate and adaptive immune responses. Cell Res. 2010;20(1):34–50.
- Weis JJ, et al. Identification of a partial cDNA clone for the C3d/Epstein-Barr virus receptor of human B lymphocytes: homology with the receptor for fragments C3b and C4b of the third and fourth components of complement. Proc Natl Acad Sci U S A. 1986;83(15):5639–5643.
- Carter RH, Fearon DT. CD19: lowering the threshold for antigen receptor stimulation of B lymphocytes. Science. 1992;256(5053):105–107.
- Barrington RA, et al. B lymphocyte memory: role of stromal cell complement and FcgammaRIIB receptors. J Exp Med. 2002;196(9):1189–1199.
- Liao F, et al. Pseudomonas aeruginosa infection induces intragraft lymphocytotoxicity that triggers lung transplant antibody-mediated rejection. Sci Transl Med. 2025;17(784):eadp1349.
- Cumpelik A, et al. Dynamic regulation of B cell complement signaling is integral to germinal center responses. Nat Immunol. 2021;22(6):757–768.
- Dempsey PW, et al. C3d of complement as a molecular adjuvant: bridging innate and acquired immunity. Science. 1996;271(5247):348–350.
- Miyamoto E, et al. Association of local intrapulmonary production of antibodies specific to donor major histocompatibility complex class I with the progression of chronic rejection of lung allografts. Transplantation. 2017;101(5):156–165.
- Miyamoto E, et al. Local intragraft humoral immune responses in chronic lung allograft dysfunction. J Heart Lung Transplant. 2025;44(1):105–117.
- Belloli EA, et al. Longitudinal forced vital capacity monitoring as a prognostic adjunct after lung transplantation. Am J Respir Crit Care Med. 2015;192(2):209–218.
- Greenland JR, et al. Bronchoalveolar lavage cell immunophenotyping facilitates diagnosis of lung allograft rejection. Am J Transplant. 2014;14(4):831–840.
- Jaramillo A, et al. Anti-HLA class I antibody binding to airway epithelial cells induces production of fibrogenic growth factors and apoptotic cell death: a possible mechanism for bronchiolitis obliterans syndrome. Hum Immunol. 2003;64(5):521–529.
- Maruyama T, et al. Induction of obliterative airway disease by anti-HLA class I antibodies. Am J Transplant. 2005;5(9):2126–2134.
- Zeng Q, et al. B cells mediate chronic allograft rejection independently of antibody production. J Clin Invest. 2014;124(3):1052–1056.
- Cumpelik A, Heeger PS. Effects of the complement system on antibody formation and function: implications for transplantation. Curr Opin Organ Transplant. 2022;27(5):399–404.
- Arcasoy SM, et al. Pathologic interpretation of transbronchial biopsy for acute rejection of lung allograft is highly variable. Am J Transplant. 2011;11(2):320–328.
- Kakoullis S, et al. Inadequate (Ax) transbronchial lung transplant biopsies-scope and the potential contributing factors. Transplant Proc. 2024;56(1):153–160.
- Levy L, et al. The impact of first untreated subclinical minimal acute rejection on risk for chronic lung allograft dysfunction or death after lung transplantation. Am J Transplant. 2020;20(1):241–249.
- Keller MB, et al. Higher molecular injury at diagnosis of acute cellular rejection increases the risk of lung allograft failure: a clinical trial. Am J Respir Crit Care Med. 2024;209(10):1238–1245.
- Yousem SA, Zeevi A. The histopathology of lung allograft dysfunction associated with the development of donor-specific HLA alloantibodies. Am J Surg Pathol. 2012;36(7):987–992.
- Agbor-Enoh S, et al. Late manifestation of alloantibody-associated injury and clinical pulmonary antibody-mediated rejection: Evidence from cell-free DNA analysis. J Heart Lung Transplant. 2018;37(7):925–932.
- Neuberger MS, Rajewsky K. Activation of mouse complement by monoclonal mouse antibodies. Eur J Immunol. 1981;11(12):1012–1016.
- Tao MH, et al. Structural features of human immunoglobulin G that determine isotype-specific differences in complement activation. J Exp Med. 1993;178(2):661–667.
- Damelang T, et al. The influence of human IgG subclass and allotype on complement activation. J Immunol. 2023;211(11):1725–1735.
- Gibson BG, et al. Contribution of animal models to the mechanistic understanding of Alternative Pathway and Amplification Loop (AP/AL)-driven Complement-mediated Diseases. Immunol Rev. 2023;313(1):194–216.
- Schaffer S, et al. Optimized isolation of renal plasma cells for flow cytometric analysis. J Immunol Methods. 2019;474:112628.
- Coppens A, et al. Murine orthotopic lung transplant models: A comprehensive overview of genetic mismatch degrees and histopathological insights into chronic lung allograft dysfunction. Am J Transplant. 2024;24(11):1930–1940.
- Lobo LJ, et al. Donor-specific antibodies are associated with antibody-mediated rejection, acute cellular rejection, bronchiolitis obliterans syndrome, and cystic fibrosis after lung transplantation. J Heart Lung Transplant. 2013;32(1):70–77.
- Safavi S, et al. De novo donor HLA-specific antibodies predict development of bronchiolitis obliterans syndrome after lung transplantation. J Heart Lung Transplant. 2014;33(12):1273–1281.
- Kauke T, et al. Bronchiolitis obliterans syndrome due to donor-specific HLA-antibodies. Tissue Antigens. 2015;86(3):178–185.
- Alnababteh M, et al. Impact of donor specific antibodies on longitudinal lung function and baseline lung allograft dysfunction. J Heart Lung Transplant. 2025;44(11):1766–1773.
- Tague LK, et al. Impact of SLCO1B3 polymorphisms on clinical outcomes in lung allograft recipients receiving mycophenolic acid. Pharmacogenomics J. 2020;20(1):69–79.
- Calabrese DR, et al. Genotypes associated with tacrolimus pharmacokinetics impact clinical outcomes in lung transplant recipients. Clin Transplant. 2018;32(8):e13332.
- Mao D, et al. Negligible role of antibodies and C5 in pregnancy loss associated exclusively with C3-dependent mechanisms through complement alternative pathway. Immunity. 2003;19(6):813–822.
- Triebwasser MP, et al. Timing and mechanism of conceptus demise in a complement regulatory membrane protein deficient mouse. Am J Reprod Immunol. 2018;80(4):e12997.
- Xu C, et al. A critical role for murine complement regulator crry in fetomaternal tolerance. Science. 2000;287(5452):498–501.
- Okazaki M, et al. A mouse model of orthotopic vascularized aerated lung transplantation. Am J Transplant. 2007;7(6):1672–1679.
- Li W, et al. Lung transplant acceptance is facilitated by early events in the graft and is associated with lymphoid neogenesis. Mucosal Immunol. 2012;5(5):544–554.
-
Version history
- Version 1 (November 11, 2025): In-Press Preview
- Version 2 (January 2, 2026): Electronic publication



Copyright © 2026 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)