Advertisement
Brief ReportImmunology Free access | 10.1172/JCI71691
T cell repertoire following autologous stem cell transplantation for multiple sclerosis
Paolo A. Muraro,1 Harlan Robins,2 Sachin Malhotra,3 Michael Howell,3 Deborah Phippard,3 Cindy Desmarais,4 Alessandra de Paula Alves Sousa,1 Linda M. Griffith,5 Noha Lim,3 Richard A. Nash,6 and Laurence A. Turka3,7
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Muraro, P. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Robins, H. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Malhotra, S. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Howell, M. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Phippard, D. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Desmarais, C. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by de Paula Alves Sousa, A. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Griffith, L. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Lim, N. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Nash, R. in: PubMed | Google Scholar
1Division of Brain Sciences, Department of Medicine, Imperial College London, London, United Kingdom. 2Fred Hutchinson Cancer Research Center, Seattle, Washington, USA. 3Immune Tolerance Network, Bethesda, Maryland, USA. 4Adaptive Biotechnologies, Seattle, Washington, USA. 5Division of Allergy, Immunology and Transplantation, National Institute of Allergy and Infectious Diseases, Bethesda, Maryland, USA. 6Colorado Blood Cancer Institute at PSL, Denver, Colorado, USA. 7Department of Surgery, Massachusetts General Hospital and Harvard Medical School, Boston, Massachusetts, USA.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Find articles by Turka, L. in: PubMed | Google Scholar
Authorship note: Paolo A. Muraro and Harlan Robins contributed equally to this work.
Published February 17, 2014 - More info
J Clin Invest. 2014;124(3):1168–1172. https://doi.org/10.1172/JCI71691.
© 2014 The American Society for Clinical Investigation
Received: June 27, 2013; Accepted: December 13, 2013
-
Abstract
Autologous hematopoietic stem cell transplantation (HSCT) is commonly employed for hematologic and non-hematologic malignancies. In clinical trials, HSCT has been evaluated for severe autoimmunity as a method to “reset” the immune system and produce a new, non-autoimmune repertoire. While the feasibility of eliminating the vast majority of mature T cells is well established, accurate and quantitative determination of the relationship of regenerated T cells to the baseline repertoire has been difficult to assess. Here, in a phase II study of HSCT for poor-prognosis multiple sclerosis, we used high-throughput deep TCRβ chain sequencing to assess millions of individual TCRs per patient sample. We found that HSCT has distinctive effects on CD4+ and CD8+ T cell repertoires. In CD4+ T cells, dominant TCR clones present before treatment were undetectable following reconstitution, and patients largely developed a new repertoire. In contrast, dominant CD8+ clones were not effectively removed, and the reconstituted CD8+ T cell repertoire was created by clonal expansion of cells present before treatment. Importantly, patients who failed to respond to treatment had less diversity in their T cell repertoire early during the reconstitution process. These results demonstrate that TCR characterization during immunomodulatory treatment is both feasible and informative, and may enable monitoring of pathogenic or protective T cell clones following HSCT and cellular therapies.
-
Introduction
When employed for immunomodulation, the goal of autologous hematopoietic stem cell transplantation (HSCT) is to create a non-autoreactive immune compartment by altering the immune repertoire and/or function of autoreactive cells (1–3). Using T cell receptor excision circle analysis in HSCT-treated MS patients, we showed previously that even in adults, thymic output contributed to the post-treatment TCR repertoire (2, 4) and that this, plus expansion of residual cells not eliminated during conditioning, regenerated the T cell compartment (5).
However, the clonal specificities of a regenerated repertoire following high-dose immunosuppressive therapy (HDIT) and autologous HSCT have not been well explored, as previous studies used CDR3 spectratyping for TCR clonal analysis, which made it impractical to sequence more than several dozen TCRs. Only a single patient receiving HSCT for autoimmune disease (ankylosing spondylitis) has had a fuller analysis of their TCR repertoire reported (6). Here, as part of a new phase II trial for poor-prognosis MS (see Methods), we have taken advantage of high-throughput deep TCRβ chain sequencing (7) to directly assess millions of TCRs per individual before and at two time points after autologous HSCT in a cohort of 24 patients.
-
Results and Discussion
As expected, HSCT rapidly eliminated most circulating T cells (Supplemental Figure 1; supplemental material available online with this article; doi: 10.1172/JCI71691DS1). Overall, neither naive nor memory CD4+ T cells recovered to baseline values by 24 months (P = 0.035 and P < 0.0001, respectively, Supplemental Figure 1, A and C). In contrast, naive CD8+ T cells recovered to baseline values by 24 months (P = 0.2184), and memory CD8+ T cells recovered to baseline by 6 months (P = 0.7754) (Supplemental Figure 1, B and D). Consistent with prior studies (8), reconstituting cells reflected a bias toward memory-phenotype cells, particularly in the CD8 lineage, where this skewing was retained even at 1–2 years after transplant, time points by which cell numbers had recovered to baseline.
We applied next-generation technology to sequence the TCRβ chains of 1 million sorted CD4+ and CD8+ T cells from each patient before transplant and 1 year after transplant (ultra-deep sequencing) as well as 200,000 CD4+ and CD8+ T cells at 2 months after transplant (deep sequencing due to lower circulating T cell numbers). Overall, from 24 enrolled patients, we obtained productive TCRβ sequence reads from nearly 100 million T cells (Supplemental Table 1).
To visualize the large quantity of data generated by sequencing, we binned the clones on the basis of their V-J usage (i.e., the combination of any one of 54 V genes with any one of 13 J genes). As shown by the example for CD4+ T cells (Supplemental Figure 2A), there was a relative dominance of a limited number of V-J combinations in the earlier (2 months) post-HSCT time point, followed by the emergence of more balanced usage at 12 months after therapy. Principal component analysis revealed that the relative V-J combination frequencies of both CD4+ and CD8+ cells clustered distinctively for each time point before and after transplant (Supplemental Figure 2B). For CD4+ T cells, the 12 month data points re-approached the baseline values, suggesting that extensive TCR skewing had begun to resolve. In contrast, in the case of CD8+ cells, there was substantial scatter of the 12 months post-HSCT samples, which were dispersed among the baseline and 2 month clusters, suggesting a high degree of interindividual variability.
To dissect the recovery of the immune repertoire after HSCT, we analyzed each clonotype identified within the libraries of TCR reads, and each clonotypic frequency was calculated as a fraction of the whole library for each sample (further details in Methods). All clones from each sample were then ranked in 4 groups according to their frequency, from 1+ or “singleton” (clones that could be assumed to have originated from a single cell) to 4+, or “top 100” clones, with one example shown in Figure 1A and a comprehensive summary for all patients shown in Figure 1B.
 Figure 1
Figure 1Proportional analysis of individual T cell clonal frequencies after HSCT. (A) Model used for classification of clonal frequency, and application of this model to a representative CD4+ and CD8+ repertoire (see text for further details). (B) Proportion of clonal classes for each participant, as defined in A, among CD4+ and CD8+ cells at baseline and at month 12.
At baseline, there was greater diversity in the CD4+ subset than in CD8+ cells, as the top 100 clones represented a lower percentage of the repertoire before transplant (median 4.93% vs. median 23.07%, respectively; P < 0.05). Importantly, HSCT led to a further restriction in diversity in both compartments. For example, at the 12 months post-transplant time point, the top 100 CD4+ and CD8 + clones now constituted a median of 9.12% and 53.61% of the T cell repertoire, respectively (both P < 0.05 vs. their baselines). Notably, in the case of CD8+ cells, the top 100 clones formed greater than 50% of the repertoire in 14 of 22 subjects, versus 0 of 22 at baseline (P < 0.05, Fisher’s exact test).
To determine how the frequency of individual clones changed within the 1+ to 4+ classification as a result of HSCT, each clone was assigned to one of 5 classes: ablated, depleted, persistent, expanded, or new, according to the criteria described in Figure 2A. We observed a striking predominance of new clones at 12 months after HSCT and reciprocal changes in the baseline clones (Figure 2A and Supplemental Figure 3). On average, the median percentage of new CD4+ clones was 81.52% in all 22 subjects (Figure 2A). The proportion of new clones was lower in CD8+ cells (Figure 2A), ranging from 12.21% to 93.21%, with a median 45.32%. Accordingly, the combined percentages of persistent and expanded clones were significantly higher in CD8+ cells than in CD4+ cells (median 54.63% vs. median 18.19%, respectively; P < 0.001).
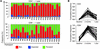 Figure 2
Figure 2Longitudinal evaluation of T cell clones after HSCT. (A) Evolution of individual clonal frequencies. “New” refers to clones not present before treatment; “expanded” clones have increased from 1+ or 2+ before treatment (see Figure 1A) to 3+ or 4+; and “persistent” clones have not changed their classification frequency. (B) Proportional representation of the clones that were found to be the 100 most frequent at 2 months after transplant in CD4+ (top) and CD8+ (bottom) T cells.
The analysis above focused on comparisons between the baseline and the 12 months post-HSCT repertoire, in part because patients were relatively lymphopenic at 2 months (Supplemental Figure 1). Nonetheless, recognizing that the cells then present would be among the first to repopulate the patients, we asked to what extent the clones observed at 2 months were present at baseline, and whether they persisted at 12 months. We found that many of the 100 most frequent CD4+ clones at 2 months, constituting 17%–71% (median 49%) of the CD4+ repertoire, were “new,” as their frequency at baseline was low to undetectable (range 0%–4%, median 0.3%; Figure 2B). At 12 months after HSCT, their frequency was reduced (range 0.1%–29%, median 4%; P < 0.05), consistent with the diversification of the repertoire. In contrast, many of the top 100 CD8+ clones seen at 2 months had expanded from clones already present at baseline, when their cumulative frequencies ranged from 1% to 27%, median 6% (Figure 2B). Of note, the top 100 CD8+ clones formed a large majority (57%–96%, median 85%) of the CD8 repertoire at 2 months after HSCT (P < 0.05 vs. baseline). Together, these data suggest that the most dominant portion of the clonal repertoire is differentially derived for CD4+ and CD8+ cells, arising de novo in the former instance, while expanding from preexisting CD8+ cells in the latter case.
To verify that alterations in the TCR repertoire observed 1 year after HSCT were the result of the transplant itself, and not merely due to repertoire “drift” over time, we examined the temporal dynamics of the TCR repertoire using paired samples of T cells taken at two time points 1, year apart, from each of 4 healthy donors. As shown for one representative example in Supplemental Figure 4, the repertoire is extremely stable, particularly when compared with that of a representative transplant patient. In 4 healthy controls, 97.1%–99.5% of the top 1,000 most frequent clones present at one sampling point remained among the top 1,000 one year later. In contrast, among all the transplanted patients, only 39.6% ± 19.8% of the top 1,000 CD4+ clones and 38.6% ± 18.9% of the top 1,000 CD8+ clones present before HSCT remained in this category 1 year after transplant (P = 0.002 for both CD4+ and CD8+ cells). Thus, the changes we observed in the TCR repertoire of transplanted patients were almost certainly due to the transplant itself, and not the 1 year interval between sample points.
Finally, we asked whether low TCR diversity (estimated using Fisher’s unseen species method; refs. 7, 9, 10), early after therapy, might be a potential measure of (re)generation of an autoimmune-biased TCR repertoire containing oligoclonal autoreactive cells. If so, then diversity would be higher in patients having a complete response (n = 19) than in those failing to meet the primary endpoint (n = 4) at the time of this interim (2 year) trial analysis. As shown in Figure 3, the repertoire reconstituted at 2 months was more diverse in responder patients (n = 19) than in the non-responder group (n = 4), and despite limited numbers of patients in the non-responder group, these differences approached statistical significance (P = 0.0694 for CD4+ cells and P = 0.0508 for CD8+ cells). This difference in diversity between responders and non-responders was lost at 1 year (P = 0.453 and 0.348 for CD4+ and CD8+ cell diversity, respectively). This suggests that TCR diversity early after transplant may be relevant to the treatment effect of HSCT, although we caution that larger patient numbers and independent follow-up studies will be needed to validate that concept.
 Figure 3
Figure 3Relationship between TCR diversity and response to HSCT. TCR diversity at 2 months after HSCT in CD4+ (top) and CD8+ (bottom) T cells divided by patients who remained stable (responder, n = 20 at baseline; and n = 19 at 2 months, as no sample was available for one patient) and who relapsed (non-responder, n = 4). The box represents the interquartile range, with the line being the median and the whiskers being 1.5× the interquartile range.
Recently, van Heijst et al. examined evolution of the TCR repertoire following allogeneic HSCT for leukemia (11). Despite differences in the technical aspects of the studies (e.g., the use of cDNA vs. genomic DNA for sequencing) and the biology of the underlying clinical situations (allogeneic vs. autologous HSCT), our studies converge in demonstrating a relationship between higher TCR diversity after HSCT and relevant clinical outcomes, i.e., low rates of infections and leukemia relapse after allogeneic HSCT and eradication of MS activity after autologous HSCT.
De novo infection and reactivation of opportunistic pathogens are complications of immunomodulatory therapy (12) and are associated with expansion of pathogen-reactive T cells. This has been demonstrated for EBV and CMV using MHC tetramers for known immunodominant epitopes in the context of common HLA types (e.g., A2 and B8). Using known public TCR amino acid sequences recognizing EBV- and CMV-derived epitopes presented by A2 or B8 (13), we observed increased clonal frequency of EBV-reactive T cells at 2 months after HSCT compared with baseline pre-transplant values in some A2 or B8 patients who experienced infection or reactivation (Supplemental Table 2; see, for example, patients P21, P3, and P14). In the most striking example, two dominant clones collectively constituted more than 35% of all CD8+ T cells in one patient (P21). Such high frequencies of EBV-reactive sequences were not observed in a recent study that used next-generation sequencing of two renal transplant recipients (14), although those patients did not receive lymphodepleting therapy. Nonetheless, changes in EBV-specific clone frequency in association with infection/activation were not uniformly observed, and EBV-specific clones were also identified in occasional patients without detectable evidence of infection (e.g., P15). This may indicate limitations in EBV diagnostics, cross-reactivity within a given TCR or TCRβ chain, our incomplete knowledge of public EBV clones, and/or the predominance of private clones in individual patients.
In summary, we found that the combination of HDIT and autologous HSCT has distinctive, and separate, effects on the reconstituting CD4+ and CD8+ repertoires. In the case of CD4+ T cells, the dominant TCR clones present before treatment were not detectable following reconstitution, and patients largely developed a new repertoire. In contrast, dominant CD8+ clones were not effectively removed, and the reconstituted CD8+ repertoire was created by clonal expansion of cells that were present before treatment. Last, and importantly, patients who failed to respond to treatment had less diversity in their T cell repertoire early during the reconstitution process, underpinning the notion that repertoire complexity is critical for the reestablishment of immune tolerance. These results demonstrate the feasibility of detailed TCR characterization during immunomodulatory therapy and investigation of the effects of HSCT on T cell repertoire diversity at the single-cell level.
-
Methods
Further information can be found in the Supplemental Methods.
TCR sequencing. Input DNA for both the pre-transplant and 1-year-post-transplant time points was 5 μg per sample, and the input DNA for the 2 months post-transplant sample was 1.2 μg. TCRβ chain sequencing was performed at Adaptive Biotechnologies using the ImmunoSEQ platform (7) with primers specific for all 54 known expressed Vβ and all 13 Jβ regions. This sequencing strategy is sufficient to achieve at least 4-fold coverage of every original template, which we have shown is sufficient to prevent a sampling effect (7). While the NKT population does have rearranged TCRs, the majority of these cells are iNKT cells, which have a highly restricted V and J usage (Vβ25 or Vβ11 in the older nomenclature). As a result, the potential contribution from these cells is capped by the very small number of TCRs with this restricted V and J subset.
Statistics. Histograms were generated to show the number of unique clones across the range of frequencies. The frequencies of each unique clone were first transformed using log10, and then the unique clones were counted within equally spaced bins. Clonotypes were grouped into 4 classes according to their frequency: 1+ denotes sequences that, based on read counts and input cell numbers, were designated as coming from single cells; 2+ refers to non-single-cell clones that were up to the 95 percentile of frequency; 3+ refers to clones between 2+ and 4+; and 4+ denotes the 100 most frequent clones within each sample.
Reader-enabled data analysis. Data and figures are accessible through the Immune Tolerance Network (ITN) TrialShare system ( https://www.itntrialshare.org/haltms.html), which allows the user to filter the underlying data and generate figures and results of analysis in addition to those submitted as part of the manuscript.
Study approval. The HALT-MS study (a Phase II Study of High-Dose Immunosuppressive Therapy [HDIT] Using Carmustine, Etoposide, Cytarabine and Melphalan [BEAM] + Thymoglobulin, and Autologous CD34+ Hematopoietic Stem Cell Transplant [HCT] for the Treatment of Poor Prognosis Multiple Sclerosis) was sponsored by NIAID and conducted by the ITN (ITN033AI) (ClinicalTrials.gov NCT00288626). The protocol was reviewed and approved by the IRB at each of the clinical sites, and all subjects provided informed consent prior to their participation. This prospective, open-label, single-arm, multicenter clinical trial was designed to determine the 5-year durability of disease stabilization in MS subjects after HDIT and autologous HCT, with an interim analysis at 2 years (15).
-
Acknowledgments
This work was supported by the ITN, funded by the NIAID, with support from the NIDDK and the Juvenile Diabetes Research Foundation (JDRF). Additional support was received from the Autoimmune Disease Clinical Trials Statistical and Clinical Coordinating Center funded by the NIAID. P.A. Muraro was supported by the UK MS society (ref. 938/10). A.P.A. Sousa was supported by the Multiple Sclerosis Trial Collaboration (ref. WMCN P36769). We thank Alice Long (Benaroya Research Institute) for outstanding technical assistance. We also thank the patients who participated in this study.
Address correspondence to: Laurence A. Turka, Transplantation Biology Research Center MGH-East, Bldg. 149-9019, 13th Street, Boston, Massachusetts 02129, USA. Phone: 617.724.7740; Fax: 617.726.6925; E-mail: lturka@partners.org.
-
Footnotes
Conflict of interest: Harlan Robins owns stock in and receives consulting fees from Adaptive Biotechnologies. Cindy Desmarais owns stock in and is an employee of Adaptive Biotechnologies. Laurence Turka owns equity in and has a family member employed by Novartis.
Reference information: J Clin Invest. 2014;124(3):1168–1172. doi:10.1172/JCI71691.
-
References
- Sykes M, Nikolic B. Treatment of severe autoimmune disease by stem-cell transplantation. Nature. 2005;435(7042):620–627.
- Muraro PA, et al. Thymic output generates a new and diverse TCR repertoire after autologous stem cell transplantation in multiple sclerosis patients. J Exp Med. 2005;201(5):805–816.
- de Kleer I, et al. Autologous stem cell transplantation for autoimmunity induces immunologic self-tolerance by reprogramming autoreactive T cells and restoring the CD4+CD25+ immune regulatory network. Blood. 2006;107(4):1696–1702.
- Muraro PA, Douek DC. Renewing the T cell repertoire to arrest autoimmune aggression. Trends Immunol. 2006;27(2):61–67.
- Dubinsky AN, Burt RK, Martin R, Muraro PA. T-cell clones persisting in the circulation after autologous hematopoietic SCT are undetectable in the peripheral CD34+ selected graft. Bone Marrow Transplant. 2010;45(2):325–331.
- Mamedov IZ, et al. Quantitative tracking of T cell clones after haematopoietic stem cell transplantation. EMBO Mol Med. 2011;3(4):201–207.
- Robins HS, et al. Comprehensive assessment of T-cell receptor beta-chain diversity in alphabeta T cells. Blood. 2009;114(19):4099–4107.
- Bosch M, Khan FM, Storek J. Immune reconstitution after hematopoietic cell transplantation. Curr Opin Hematol. 2012;19(4):324–335.
- Efron B, Thisted R. Estimating the number of unseen species: how many words did Shakespeare know? Biometrika. 1976;63(3):435–447.
- Fisher RA, Corbet AS, Williams CB. The relation between the number of species and the number of individuals in a random sample of an animal population. J Anim Ecol. 1943;12(1):42–58.View this article via: CrossRef Google Scholar
- van Heijst JW, et al. Quantitative assessment of T cell repertoire recovery after hematopoietic stem cell transplantation. Nat Med. 2013;19(3):372–377.
- Fishman JA. Infection in solid-organ transplant recipients. N Engl J Med. 2007;357(25):2601–2614.
- Miles JJ, Douek DC, Price DA. Bias in the alphabeta T-cell repertoire: implications for disease pathogenesis and vaccination. Immunol Cell Biol. 2011;89(3):375–387.
- Klarenbeek PL, et al. Deep sequencing of antiviral T-cell responses to HCMV and EBV in humans reveals a stable repertoire that is maintained for many years. PLoS Pathog. 2012;8(9):e1002889.
- Nash RA, et al. Treatment of severe relapsing-remitting multiple sclerosis with high-dose immunosuppressive therapy and autologous hematopoietic cell transplantation: 2-year follow-up results of the HALT MS Clinical Trial (Immune Tolerance Network: ITN033AI). Biol Blood Marrow Transplant. 2013;19(2 suppl):S129.
-
Version history
- Version 1 (February 17, 2014): No description
- Version 2 (March 3, 2014): No description



Copyright © 2026 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)



