Advertisement
Commentary Free access | 10.1172/JCI59219
Parkin reinvents itself to regulate fatty acid metabolism by tagging CD36
Nada A. Abumrad1 and Darren J. Moore2
1Washington University School of Medicine, Department of Medicine, Center for Human Nutrition, St. Louis, Missouri, USA. 2Ecole Polytechnique Fédérale de Lausanne (EPFL), Brain Mind Institute, School of Life Sciences, Lausanne, Switzerland.
Address correspondence to: Nada A. Abumrad, Department of Medicine, Campus Box 8031, Washington University School of Medicine, St. Louis, Missouri 63110, USA. Phone: 314.747.0348; Fax: 314.444.3432; E-mail: nabumrad@wustl.edu.
Find articles by Abumrad, N. in: JCI | PubMed | Google Scholar
1Washington University School of Medicine, Department of Medicine, Center for Human Nutrition, St. Louis, Missouri, USA. 2Ecole Polytechnique Fédérale de Lausanne (EPFL), Brain Mind Institute, School of Life Sciences, Lausanne, Switzerland.
Address correspondence to: Nada A. Abumrad, Department of Medicine, Campus Box 8031, Washington University School of Medicine, St. Louis, Missouri 63110, USA. Phone: 314.747.0348; Fax: 314.444.3432; E-mail: nabumrad@wustl.edu.
Find articles by Moore, D. in: JCI | PubMed | Google Scholar
Published August 25, 2011 - More info
J Clin Invest. 2011;121(9):3389–3392. https://doi.org/10.1172/JCI59219.
© 2011 The American Society for Clinical Investigation
Related article:
Abstract
It has long been hypothesized that abnormalities in lipid biology contribute to degenerative brain diseases. Consistent with this, emerging epidemiologic evidence links lipid alterations with Parkinson disease (PD), and disruption of lipid metabolism has been found to predispose to α-synuclein toxicity. We therefore investigated whether Parkin, an E3 ubiquitin ligase found to be defective in patients with early onset PD, regulates systemic lipid metabolism. We perturbed lipid levels by exposing Parkin+/+ and Parkin–/– mice to a high-fat and -cholesterol diet (HFD). Parkin–/– mice resisted weight gain, steatohepatitis, and insulin resistance. In wild-type mice, the HFD markedly increased hepatic Parkin levels in parallel with lipid transport proteins, including CD36, Sr-B1, and FABP. These lipid transport proteins were not induced in Parkin–/– mice. The role of Parkin in fat uptake was confirmed by increased oleate accumulation in hepatocytes overexpressing Parkin and decreased uptake in Parkin–/– mouse embryonic fibroblasts and patient cells harboring complex heterozygous mutations in the Parkin-encoding gene PARK2. Parkin conferred this effect, in part, via ubiquitin-mediated stabilization of the lipid transporter CD36. Reconstitution of Parkin restored hepatic fat uptake and CD36 levels in Parkin–/– mice, and Parkin augmented fat accumulation during adipocyte differentiation. These results demonstrate that Parkin is regulated in a lipid-dependent manner and modulates systemic fat uptake via ubiquitin ligase–dependent effects. Whether this metabolic regulation contributes to premature Parkinsonism warrants investigation.
Authors
Kye-Young Kim, Mark V. Stevens, M. Hasina Akter, Sarah E. Rusk, Robert J. Huang, Alexandra Cohen, Audrey Noguchi, Danielle Springer, Alexander V. Bocharov, Tomas L. Eggerman, Der-Fen Suen, Richard J. Youle, Marcelo Amar, Alan T. Remaley, Michael N. Sack
-
Abstract
Parkinson disease (PD) is a relatively common neurodegenerative disorder characterized by the progressive degeneration of dopaminergic neurons in the substantia nigra. About 5%–10% of PD cases are inherited. Mutations in the Parkin gene, which encodes a protein that can function as an E3 ubiquitin ligase, are a common cause of familial PD. Such mutations act in a loss-of-function manner and impair the ability of the encoded protein to mediate substrate ubiquitination, although the subsequent molecular pathway that precipitates neuronal degeneration is poorly defined. In this issue of the JCI, Kim and colleagues describe painstaking evidence using a number of dissecting approaches in intact animals and cultured cells to functionally link Parkin and the class B scavenger receptor CD36, suggesting a novel and complex connection between PD and fatty acid metabolism.
Parkinson disease (PD) affects 1%–2% of the world’s population over the age of 65 years. It is more common in developed countries where people live longer. PD is characterized by the progressive degeneration of dopaminergic neurons in the substantia nigra, which leads to diminished striatal dopamine levels and the appearance of intracytoplasmic proteinaceous inclusions, termed Lewy bodies, in surviving brainstem neurons. The cardinal motor symptoms of PD include bradykinesia, resting tremor, muscular rigidity, and postural instability. There are also nonmotor symptoms, which include autonomic, cognitive, and psychiatric disturbances. About 5%–10% of PD cases are inherited, with at least 16 chromosomal loci (PARK1–16) associated with these familial forms of disease (1). Mutations in the α-synuclein gene (SNCA, which falls into both the PARK1 and PARK4 loci) and the leucine-rich repeat kinase 2 gene (LRRK2, which accounts for the PARK8 locus) unambiguously cause autosomal dominant PD, whereas mutations in the Parkin gene (which accounts for the PARK2 locus), the PTEN-induced putative kinase 1 gene (PINK1, which accounts for the PARK6 locus), the DJ-1 gene (which accounts for the PARK7 locus), and the ATPase type 13A2 gene (ATP13A2, which accounts for the PARK9 locus) cause autosomal recessive forms of the disease (1). Idiopathic PD is considered to result from a combination of genetic susceptibility, aging, and environmental factors (for example, pesticides) (2).
-
Parkin: a ubiquitin ligase linked to PD
Mutations in the Parkin gene on chromosome 6 cause autosomal recessive, juvenile-onset parkinsonism (AR-JP), the most common form of early-onset disease (3, 4). The Parkin gene encodes a 465 amino acid protein consisting of a ubiquitin-like domain and a RING-box domain containing two RING finger motifs separated by an in-between-RING finger motif (Figure 1 and ref. 5). Similar to other RING-box proteins, Parkin can function as an E3 ubiquitin ligase to catalyze, in cooperation with E1-activating and E2-conjugating enzymes, the covalent attachment of the carboxyl terminal glycine residue of ubiquitin to an internal lysine residue of protein substrates (5). Parkin can link substrates to chains of ubiquitin polymerized through different lysine residues of the ubiquitin components, with K48-linked chains targeting substrates for proteasomal degradation and K63-linked chains acting as a nondegradative signal (5). Parkin can also mediate monoubiquitination of proteins and itself (6). Familial mutations in Parkin act in a loss-of-function manner and impair substrate ubiquitination, although the subsequent molecular pathway that precipitates neuronal degeneration is poorly defined. In this issue of the JCI, Kim et al. have identified a new substrate of Parkin-mediated ubiquitination that provides a novel and complex connection between fatty acid (FA) metabolism and PD (7).
 Figure 1
Figure 1Substrates of Parkin-dependent ubiquitination. Parkin contains a ubiquitin-like (Ubl) domain and a RING-box (RBR) domain consisting of two RING finger motifs separated by an in-between-RING finger domain (IBR). Parkin cooperates with E1-activating and E2-conjugating enzymes to covalently modify protein substrates with ubiquitin, either via alternatively linked polyubiquitin chains or via monoubiquitination. Polyubiquitination via K48-linked chains serves to target substrates for proteasomal degradation, whereas K63-linked polyubiquitination and monoubiquitination serve as nondegradative signals. A broad range of substrates is ubiquitinated by Parkin, often being targeted for degradation (i.e., PAEL-R, AIMP2/p38, CDCrel-1, and cyclin E), whereas others are degradation independent (i.e., Hsp70 and synphilin-1). The lipid transporter CD36 is the latest substrate identified for Parkin and is stabilized by monoubiquitination (7), providing a novel mechanism for the regulation of FA uptake.
-
Parkin finds a new substrate
A number of substrates have been identified for Parkin (Figure 1), including numerous proteins of the outer mitochondrial membrane (5, 8). However, impaired ubiquitination of these has yet to be linked molecularly to PD. The study by Kim et al. (7) identifies the class B scavenger receptor CD36 (also known as FA translocase [FAT]) as a new substrate of Parkin-mediated ubiquitination and proposes a potentially broad function of Parkin in the regulation of FA metabolism. CD36 is a 472–amino acid transmembrane protein that binds with high affinity to a number of lipid ligands including long chain FAs, anionic phospholipids, and native or modified lipoproteins. CD36 also recognizes a number of nonlipid ligands such as thrombospondin-1, collagen, and amyloid-β. Interaction of CD36 with its ligands elicits a range of intracellular signaling processes involving Src and MAP kinases that integrate lipid metabolism and inflammation and also involve CD36 in pathways related to oxidative stress, angiogenesis, platelet hyperactivity, phagocytosis, and cell migration (9, 10). Studies in CD36-deficient mice (11) and humans (12) have linked CD36 to FA and glucose utilization. Common polymorphisms in the CD36 gene contribute to individual variability in blood lipids (13) and to differences in platelet function (14). They may also influence susceptibility to the metabolic syndrome (13) and diabetes (15). Overall, although CD36 is clearly multifunctional, data currently available in humans mainly implicate the protein in FA and lipoprotein metabolism and associated abnormalities.
The regulation of CD36 by Parkin uncovered by Kim et al. (7) is consistent with epidemiologic studies linking altered lipid metabolism to PD (16, 17) and with evidence that abnormalities in lipid processing contribute to degenerative brain diseases (18). Indeed, these earlier findings are what led Kim and colleagues to investigate the possibility that Parkin regulates systemic lipid metabolism by characterizing the global metabolic responses of WT and Parkin-deficient (Parkin–/–) mice to high-fat diet (HFD) (7). As expected, the WT mice gained weight on the diet and developed insulin resistance and lipid accumulation in the liver, which was associated with parallel increases in lipid transport proteins and Parkin. In contrast, the Parkin–/– mice were resistant to all these metabolic changes. CD36 was among the lipid transport proteins altered by HFD in the WT but not Parkin–/– mice, and CD36 levels paralleled those of Parkin in multiple tissues. Importantly, rescue of CD36 expression in the liver of Parkin–/– mice increased hepatic fat uptake while rescue of Parkin expression restored endogenous CD36 levels and lipid accumulation. Gain- and loss-of-function studies in cultured cells documented that Parkin regulates CD36 levels and FA uptake, while immunoprecipitation experiments suggested a physical interaction between CD36 and Parkin. Reduction of FA uptake was also observed in B cells derived from patients harboring heterozygous Parkin mutations that associate with early-onset parkinsonism.
Kim et al. (7) went on to show that the regulation of CD36 by Parkin is mediated via its monoubiquitination, which they propose stabilizes CD36 at the plasma membrane. This contrasts with FAs, which were previously shown to induce polyubiquitination of CD36 and target it for proteasomal degradation. The lysine residues within CD36 that are ubiquitinated upon exposure to FAs are K469 and K472 in the carboxyl terminus of the protein (10). One of these residues is likely targeted by Parkin, since these are the only lysines present in the cytoplasmic domains of CD36. These residues could constitute a site where the influences of FAs and Parkin on CD36 are integrated. Consistent with FAs and Parkin competing to regulate CD36 levels, Kim et al. (7) found that Parkin prevents FA-mediated reduction in CD36 levels. FA induction of CD36 polyubiquitination and degradation acts to limit cellular toxicity from high FAs and would be blunted in the presence of Parkin. Whether this means that Parkin contributes to abnormalities of FA utilization under conditions of high FAs remains to be determined. However, it is clear from the data generated by Kim et al. (7) that Parkin is a player in the regulation of lipid metabolism, especially considering that its expression is not limited to the brain and is observed in a number of metabolically active tissues including the heart and skeletal muscle, where it is particularly abundant. This interpretation would be consistent with previous reports of a role for Parkin in oxidative metabolism and in maintaining mitochondrial quality control. Identified first in Drosophila, where it was reported that Parkin-null flies cannot climb or fly and exhibit reduced body mass (19), it was later shown in various mammalian cell types that under stressful conditions, Parkin selectively translocates to damaged mitochondria to ubiquitinate mitochondrial proteins and induce mitophagy (8). It is worth noting that the Parkin substrate CD36 has been identified in isolated mitochondrial preparations, especially under states of high-energy demand such as exercise, where it would facilitate FA access to metabolizing enzymes (20).
-
FAs and PD
The paper by Kim et al. (7) shows that Parkin is lipid sensitive and is regulated by dietary fat intake. So, does FA metabolism modulate susceptibility to PD? Emerging epidemiological evidence lends some support to this. In particular, increased dietary intake of omega-3 polyunsaturated FAs (n-3 PUFAs) has been associated with a decreased risk of PD (16), while increased PD risk has been associated with a high intake of the n-6 PUFA arachidonic acid (21). Dietary n-3 PUFAs also provide neuroprotection in toxin-induced rodent models of parkinsonism (16). These studies suggest a role for FA metabolism in the maintenance and survival of nigrostriatal pathway dopaminergic neurons. They also touch on the importance of arachidonic acid metabolism in the brain and how its dysregulation may associate with disease (22). Arachidonic acid, released from membrane phospholipid by phospholipases, notably cytosolic phospholipase A2 (cPLA2), and its metabolites are involved in the regulation of synaptic transmission, and this may be modulated by the PD-associated protein α-synuclein, which sequesters the released arachidonic acid (23). Coincidentally, the novel Parkin substrate CD36, identified by Kim et al. (7), is also involved in the regulation of arachidonic acid levels. CD36 facilitates arachidonic acid release by virtue of its role in activating membrane calcium channels and cPLA2 (24). It is tempting therefore to speculate that Parkin, CD36, and α-synuclein (Figure 2) may interact to control arachidonic acid metabolism in the brain. It is known that excess arachidonic acid release by activated brain microglia results in the formation of neurotoxic compounds such as prostaglandins, cytokines, and reactive oxygen species (18). Misfolded or aggregated α-synuclein (like amyloid-β) would potentiate these events. Studies using transgenic mice or cultured primary microglia overexpressing human α-synuclein mutations showed early microglial activation that was partially inhibited in microglia derived from CD36-deficient mice (25).
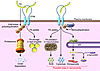 Figure 2
Figure 2Potential functional interactions among Parkin, CD36, and the PD-associated protein α-synuclein. Parkin monoubiquitinates CD36, stabilizing it and increasing its level at the plasma membrane. This is in contrast to long-chain FAs, which induce CD36 polyubiquitination and target CD36 for proteosomal degradation. At the plasma membrane, CD36 facilitates uptake of FAs, which are stored into lipid droplets or oxidized by mitochondria. Under conditions of cellular stress, CD36 is involved in Ca2+- and ERK1/2-mediated activation of cPLA2 to release arachidonic acid (AA) from endoplasmic reticulum and nuclear membrane phospholipids. Brain AA is involved in the regulation of synaptic transmission and is also converted by microglia to bioactive eicosanoids, which can have pleiotropic effects on inflammation. Excess production of AA-derived metabolites, ROS (from mitochondria), and cytokines (CD36 signaling contributes to cytokine production) by the activated microglia can lead to neurotoxicity. Binding of AA by α-synuclein could modulate its effects and its metabolic conversion to eicosanoids.
In summary, the paper by Kim et al. (7) brings novel insight into the well-known connection between neurodegenerative disorders and lipid metabolism. By documenting the clear link between Parkin and CD36, it introduces a new area of research with abundant possibilities. In addition, the paper shows that Parkin levels are regulated by lipid, an important finding that provides a concrete avenue by which dietary fat intake can have an impact on PD etiology. Possibly, we should ask, does aging increase PD susceptibility via the associated alterations in lipid metabolism?
-
Acknowledgments
Work in the lab of N.A. Abumrad related to CD36 ubiquitination by FAs and its role in arachidonic acid release is supported by NIHI grants P01 HL057278, DK33301, and DK56351.
Address correspondence to: Nada A. Abumrad, Department of Medicine, Campus Box 8031, Washington University School of Medicine, St. Louis, Missouri 63110, USA. Phone: 314.747.0348; Fax: 314.444.3432; E-mail: nabumrad@wustl.edu.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Reference information: J Clin Invest. 2011;121(9):3389–3392. doi:10.1172/JCI59219
See the related article at Parkin is a lipid-responsive regulator of fat uptake in mice and mutant human cells.
-
References
- Gasser T. Mendelian forms of Parkinson’s disease. Biochim Biophys Acta. 2009;1792(7):587–596.View this article via: PubMed Google Scholar
- Wirdefeldt K, Adami HO, Cole P, Trichopoulos D, Mandel J. Epidemiology and etiology of Parkinson’s disease: a review of the evidence. Eur J Epidemiol. 2011;26(suppl 1):S1–S58.View this article via: PubMed Google Scholar
- Kitada T, et al. Mutations in the Parkin gene cause autosomal recessive juvenile parkinsonism. Nature. 1998;392(6676):605–608.
- West AB, Maidment NT. Genetics of Parkin-linked disease. Hum Genet. 2004;114(4):327–336.
- Moore DJ. Parkin: a multifaceted ubiquitin ligase. Biochem Soc Trans. 2006;34(pt 5):749–753.
- Moore DJ, West AB, Dikeman DA, Dawson VL, Dawson TM. Parkin mediates the degradation-independent ubiquitination of Hsp70. J Neurochem. 2008;105(5):1806–1819.
- Kim K-Y, et al. Parkin is a lipid-responsive regulator of fat uptake in mice and mutant human cells. J Clin Invest. 2011;121(9):3701–3712.View this article via: JCI Google Scholar
- Chan NC, et al. Broad activation of the ubiquitin-proteasome system by Parkin is critical for mitophagy. Hum Mol Genet. 2011;20(9):1726–1737.
- Silverstein RL, Febbraio M. CD36, a scavenger receptor involved in immunity, metabolism, angiogenesis, and behavior. Sci Signal. 2009;2(72):re3.
- Su X, Abumrad NA. Cellular fatty acid uptake: a pathway under construction. Trends Endocrinol Metab. 2009;20(2):72–77.
- Hajri T, Han XX, Bonen A, Abumrad NA. Defective fatty acid uptake modulates insulin responsiveness and metabolic responses to diet in CD36-null mice. J Clin Invest. 2002;109(10):1381–1389.
- Yamashita S, et al. Physiological and pathological roles of a multi-ligand receptor CD36 in atherogenesis; insights from CD36-deficient patients. Mol Cell Biochem. 2007;299(1–2):19–22.
- Love-Gregory L, et al. Common CD36 SNPs reduce protein expression and may contribute to a protective atherogenic profile. Hum Mol Genet. 2011;20(1):193–201.View this article via: PubMed Google Scholar
- Ghosh A, et al. Platelet CD36 surface expression levels affect functional responses to oxidized LDL and are associated with inheritance of specific genetic polymorphisms. Blood. 2011;117(23):6355–6366.
- Lepretre F, et al. A CD36 nonsense mutation associated with insulin resistance and familial type 2 diabetes. Hum Mutat. 2004;24(1):104.View this article via: PubMed Google Scholar
- Bousquet M, Calon F, Cicchetti F. Impact of omega-3 fatty acids in Parkinson’s disease [published online ahead of print March 23, 2011]. Ageing Res Rev . doi:10.1016/j.arr.2011.03.001.
- de Lau LM, Bornebroek M, Witteman JC, Hofman A, Koudstaal PJ, Breteler MM. Dietary fatty acids and the risk of Parkinson disease: the Rotterdam study. Neurology. 2005;64(12):2040–2045.
- Farooqui T, Farooqui AA. Lipid-mediated oxidative stress and inflammation in the pathogenesis of Parkinson’s disease. Parkinsons Dis. 2011;2011:247467.View this article via: PubMed Google Scholar
- Greene JC, Whitworth AJ, Kuo I, Andrews LA, Feany MB, Pallanck LJ. Mitochondrial pathology and apoptotic muscle degeneration in Drosophila Parkin mutants. Proc Natl Acad Sci U S A. 2003;100(7):4078–4083.View this article via: PubMed Google Scholar
- Smith BK, et al. FAT/CD36 is located on the outer mitochondrial membrane, upstream of long-chain acyl-CoA synthetase, and regulates palmitate oxidation. Biochem J. 2011;437(1):125–134.
- Miyake Y, et al. Dietary fat intake and risk of Parkinson’s disease: a case-control study in Japan. J Neurol Sci. 2010;288(1–2):117–122.
- Farooqui AA, Horrocks LA, Farooqui T. Interactions between neural membrane glycerophospholipid and sphingolipid mediators: a recipe for neural cell survival or suicide. J Neurosci Res. 2007;85(9):1834–1850.View this article via: PubMed Google Scholar
- Darios F, Ruiperez V, Lopez I, Villanueva J, Gutierrez LM, Davletov B. Alpha-synuclein sequesters arachidonic acid to modulate SNARE-mediated exocytosis. EMBO Rep. 2011;11(7):528–533.
- Kuda O, et al. CD36 Protein Is Involved in Store-operated Calcium Flux, Phospholipase A2 Activation, and Production of Prostaglandin E2. J Biol Chem. 2011;286(20):17785–17795.
- Su X, Federoff HJ, Maguire-Zeiss KA. Mutant alpha-synuclein overexpression mediates early proinflammatory activity. Neurotox Res. 2009;16(3):238–254.
- Gasser T. Mendelian forms of Parkinson’s disease. Biochim Biophys Acta. 2009;1792(7):587–596.
-
Version history
- Version 1 (August 25, 2011): No description
- Version 2 (September 1, 2011): No description



Copyright © 2024 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)


