Advertisement
Brief ReportImmunology Free access | 10.1172/JCI44254
A leaky mutation in CD3D differentially affects αβ and γδ T cells and leads to a Tαβ–Tγδ+B+NK+ human SCID
Juana Gil,1 Elena M. Busto,2 Beatriz Garcillán,2 Carmen Chean,1 Maria Cruz García-Rodríguez,3 Andrea Díaz-Alderete,1 Joaquín Navarro,1 Jesús Reiné,2 Angeles Mencía,4 Dolores Gurbindo,1 Cristina Beléndez,1 Isabel Gordillo,1 Marlena Duchniewicz,5 Kerstin Höhne,5 Félix García-Sánchez,6 Eduardo Fernández-Cruz,1 Eduardo López-Granados,3 Wolfgang W.A. Schamel,5 Miguel A. Moreno-Pelayo,4 María J. Recio,2 and José R. Regueiro2
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Gil, J. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Busto, E. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Garcillán, B. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Chean, C. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by García-Rodríguez, M. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Díaz-Alderete, A. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Navarro, J. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Reiné, J. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Mencía, A. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Gurbindo, D. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Beléndez, C. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Gordillo, I. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Duchniewicz, M. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Höhne, K. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by García-Sánchez, F. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Fernández-Cruz, E. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by López-Granados, E. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Schamel, W. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Moreno-Pelayo, M. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Recio, M. in: PubMed | Google Scholar
1Gregorio Marañón University Hospital, Madrid, Spain. 2School of Medicine, Complutense University, i+12 and RIER, Madrid, Spain. 3La Paz University Hospital, Madrid, Spain. 4Ramón y Cajal University Hospital, IRYCIS and CIBERER, Madrid, Spain. 5Faculty of Biology, Centre of Chronic Immunodeficiency (CCI), BIOSS Centre for Biological Signaling Studies, University of Freiburg and Max Planck Institute of Immunobiology and Epigenetics, Freiburg, Germany. 6Blood Transfusion Centre, Madrid, Spain.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Find articles by Regueiro, J. in: PubMed | Google Scholar
Authorship note: Juana Gil and Elena M. Busto are co–first authors. María J. Recio and José R. Regueiro are co–senior authors.
Published September 19, 2011 - More info
J Clin Invest. 2011;121(10):3872–3876. https://doi.org/10.1172/JCI44254.
© 2011 The American Society for Clinical Investigation
Received: January 18, 2011; Accepted: August 3, 2011
-
Abstract
T cells recognize antigens via their cell surface TCR and are classified as either αβ or γδ depending on the variable chains in their TCR, α and β or γ and δ, respectively. Both αβ and γδ TCRs also contain several invariant chains, including CD3δ, which support surface TCR expression and transduce the TCR signal. Mutations in variable chains would be expected to affect a single T cell lineage, while mutations in the invariant chains would affect all T cells. Consistent with this, all CD3δ-deficient patients described to date showed a complete block in T cell development. However, CD3δ-KO mice have an αβ T cell–specific defect. Here, we report 2 unrelated cases of SCID with a selective block in αβ but not in γδ T cell development, associated with a new splicing mutation in the CD3D gene. The patients’ T cells showed reduced CD3D transcripts, CD3δ proteins, surface TCR, and early TCR signaling. Their lymph nodes showed severe T cell depletion, recent thymus emigrants in peripheral blood were strongly decreased, and the scant αβ T cells were oligoclonal. T cell–dependent B cell functions were also impaired, despite the presence of normal B cell numbers. Strikingly, despite the specific loss of αβ T cells, surface TCR expression was more reduced in γδ than in αβ T cells. Analysis of individuals with this CD3D mutation thus demonstrates the contrasting CD3δ requirements for αβ versus γδ T cell development and TCR expression in humans and highlights the diagnostic and clinical relevance of studying both TCR isotypes when a T cell defect is suspected.
-
Introduction
T lymphocytes recognize antigens by means of a cell surface complex termed the TCR. The TCR contains 2 variable chains to bind antigens and several invariant chains to support variable chains and to transduce the signals required for T cell differentiation and antigen recognition (1). The invariant chains present in the human TCR are CD3γ, CD3δ, CD3ε, and TCRζ (CD247) (Figure 1A). T lymphocytes belong to either the αβ or the γδ lineage according to the types of variable chains incorporated into their TCR, α and β or γ and δ, respectively.
 Figure 1
Figure 1CD3D mutation analysis. (A) Structure of the 2 TCR that incorporate the CD3δ chain (arrows). EC, extracellular region; TM, transmembrane region; Cyto, cytoplasmic region. (B) CD3D RNA RT-PCR amplification products. ΔEx2 bands lack exon 2 (C) CD3δ protein and gene structure with localization of the G→A mutation. SP, signal peptide. (D) CD3D electropherograms showing the IVS2+5G→A mutation (arrow) in patient AIII.1 (left) and his father (right), and the BsaAI restriction site. Exon/intron sequences are in upper/lower case, respectively. (E) Genetic pedigrees. Circles indicate females; squares indicate males (slashes indicate deceased). Solid symbols denote homozygosity; half-solid symbols denote heterozygosity. (F) WT and ΔEx2 CD3D transcript levels relative to CD3E by quantitative RT-PCR in PBMCs using exon-specific primers and isoform-specific probes. Data represent mean ± SD of at least 2 experiments, relative to the highest value in each data set, which is shown as 1.
Mutations in TCR or CD3 genes selectively impair T cell development (2). Unless hematopoietic stem cells are replaced, the mutations frequently cause early-onset SCID and death. When a variable chain such as TCRα is affected, only αβ T cells are impaired, as shown recently (3). When an invariant chain is affected, both αβ and γδ T cells are either absent, as observed in CD3δ or CD3ε deficiency (4, 5), or reduced, as reported for CD3γ or TCRζ deficiency (6, 7). These immunophenotypes are described as T–B+NK+ or T±B+NK+, respectively. However, no selective αβ or γδ T cell–deficient patients have been reported for invariant TCR chain deficiencies. We describe 2 SCID cases with severe selective αβ T lymphopenia (Tαβ–Tγδ+B+NK+) caused by a leaky mutation in CD3D that titrated the differential CD3δ requirements for human αβ and γδ T lymphocyte development and TCR expression in vivo.
-
Results and Discussion
Case reports. Two unrelated children from nonconsanguineous Ecuadorian parents showed common clinical and immunophenotypic features. They presented at 13 (AIII.1) and 5 (BII.2) months of age with SCID features, T±B+NK+ phenotype, low CD3 expression, strongly impaired proliferative responses to T cell mitogens (Table 1), severe lymph node T cell depletion, and lack of activated germinal centers. Both received conditioning and haploidentical CD34+ peripheral blood hematopoietic stem cell transplantation at 23 and 8 months, respectively.
Patient AIII.1 was admitted with failure to thrive, bronchopneumonia, severe diarrhea caused by strains of Salmonella, Campylobacter, and Cryptosporidium, oral candidiasis, and atopic dermatitis. CMV and EBV were negative by PCR. Neutrophil, lymphocyte, and platelet numbers and serum and urine biochemistry were normal, including Ig levels except IgE, which was strongly increased (Table 1). NK cell function was normal, whereas T cell–dependent B cell function was impaired. Specific Abs against protein antigens were not induced upon vaccination and were not detected against common pathogens or autoantigens, whereas natural (IgM) Abs against polysaccharides such as isoagglutinins and heterophile Abs were normal. The patient required total parenteral nutrition, specific antimicrobials, and i.v. Ig therapy. Absence epilepsy developed at 16 months. Sclerosing cholangitis was observed before receiving a maternal transplant. At 4 years of age, he is doing well, with mixed chimerism.
Patient BII.2 was admitted with fever, prostrating diarrhea, and respiratory distress. Urine CMV and nasal adenovirus, discrete lymphopenia, and severe hypogammaglobulinemia were observed. A protein-losing enteropathy was diagnosed, and he started on i.v. Ig and prophylactic Septrin, but developed several lymphadenopathies. After an initial improvement, respiratory function deteriorated and bronchoalveolar CMV was identified, which required antivirals. He underwent paternal transplantation with full chimerism in 3 weeks, but returned to the intensive care unit with respiratory distress, hemodynamic instability, encephalopathy, and liver failure with secondary coagulopathy, and died following multiorganic failure. Necropsy evidenced a rudimentary thymus with conserved reticular structure but complete depletion of lymphocytes and Hassall corpuscles. CMV identified in the liver, brain, heart, and lung was the likely cause of decease.
A novel mutation in CD3D. The decreased CD3 expression observed in both patients suggested a potential TCR defect. We thus analyzed CD3 and CD247 RNA by RT-PCR and found short CD3D PCR products in several family members (Figure 1B). Sequencing revealed a complete in-frame deletion of exon 2, which encodes the extracellular Ig-like domain of CD3δ (Figure 1C).
Exon skipping suggested a potential splicing defect. Genomic DNA sequencing detected a homozygous G-to-A mutation at position +5 in the 5′ splice donor site of intron 2 (IVS2+5G→A; Figure 1D). The mutation abrogated a restriction site for the enzyme BsaAI, which was used to follow its segregation (Figure 1E). The mutation was causing the immunodeficiency, as it was not found in 140 Spanish or Ecuadorian healthy donors, and its location is strictly conserved in mammals (Supplemental Figure 1; supplemental material available online with this article; doi: 10.1172/JCI44254DS1).
The patients’ parents were carriers of the same CD3D mutation and had a similar geographic origin, indicating that they likely shared a founder mutant allele. The analysis of polymorphic microsatellite markers spanning the CD3 region (which contains CD3G, CD3D, and CD3E) confirmed the presence of a shared core haplotype associated with the mutation (Supplemental Figure 2).
From the data in Figure 1B, it seemed that some normal PCR products might be present in the patients, indicating that the mutation did not abrogate normal splicing at the CD3D locus, as shown in similar intronic mutations (8). To analyze CD3D splicing, quantitative RT-PCR relative to CD3E was performed. The results confirmed the presence of small amounts of WT CD3D transcripts in the patients (Figure 1F).
From these results, we concluded that a homozygous IVS2+5G→A mutation strongly impaired (around 10-fold), but did not abrogate, normal CD3D splicing in both patients.
Reduced CD3δ protein. The small levels of WT CD3D transcripts were found to be sufficient to encode for half-normal levels of WT CD3δ proteins in the patients’ T cells as shown in family B by Western blotting (Figure 2) or intracellular flow cytometry (Supplemental Figure 3A). In contrast, the dominant ΔEx2 CD3D transcripts did not give rise to detectable levels of the predicted headless CD3δ chain, despite being readily detected after transfection in non-T cells (Figure 2B).
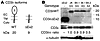 Figure 2
Figure 2CD3δ protein analysis. (A) WT CD3δ and the predicted truncated isoform (ΔEx2). (B) Western blotting analysis of CD3δ isoforms in lysates from ΔEx2-transfected 293T cells, Jurkat cells (JK), or T cell lines with the indicated CD3D IVS2+5 genotypes using APA1/2 (anti-human CD3δ cytoplasmic tail), anti-CD3ε, or anti-tubulin. The numbers under each lane indicate CD3δWT band intensity relative to CD3ε.
We cannot exclude that small amounts of ΔEx2 CD3δ might be expressed below the detection limit of Western blotting. However, when overexpressed in Drosophila cells, ΔEx2 did not compete with WT CD3δ in the formation of a TCR complex (Supplemental Methods and Supplemental Figure 4).
From these studies, we concluded that the immunodeficiency was associated with reduced levels of normal CD3δ.
Immunological characteristics. The human CD3δ chain is incorporated into both TCRαβ and TCRγδ (Figure 1A). To establish how the CD3D mutation affected lymphocyte differentiation, the numbers of αβ T cells and γδ T cells were determined in both patients (Figure 3A). The results showed a severe selective reduction in peripheral blood αβ T lymphocyte numbers (both CD4+ and CD8+; Supplemental Figure 5), close to 10-fold compared with the median value of healthy age-matched controls. In contrast, γδ T cells as well as B and NK lymphocytes were detected in normal numbers (Tαβ–Tγδ+B+NK+ phenotype; Figure 3A and Table 1). A possible role for CMV in γδ T cell predominance was proposed in some SCID reports (9) but not in others (10). However, CMV-induced γδ T expansion was excluded in patient AIII.1.
 Figure 3
Figure 3T lymphocyte analysis. (A) Absolute cell numbers in patients plotted as a function of age in comparison with the normal age-matched distribution (P5, P50, and P95). The leukocyte subsets are lymphocytes (Ly), eosinophils (Eo), αβ T (CD4++CD8bright), and γδ T (11F2+) cells. (B) TCRαβ and TCRγδ cell surface expression in primary lymphocytes from the patients (dashed lines, AA genotype) in comparison with controls (solid lines, GG genotype), measured using the indicated TCRαβ- and TCRγδ-specific mAb. The numbers in each histogram indicate MFI ratios between control and patient. (C) CD69 induction (geometric MFI increments) after 24 hours in anti-CD3–stimulated (dashed lines) versus unstimulated (solid lines) primary αβ T (CD4+) or γδ T (IMMU510+) lymphocytes with the indicated genotypes. (D) Patient T cell growth in feeder cell cultures expressed as a percentage of carrier BI.1 T cell numbers.
The reduction caused by the CD3D mutation in αβ, but not γδ, T lymphocyte numbers suggested a differential CD3δ requirement for TCR expression or function in αβ versus γδ T cells. Counterintuitively, TCR expression was around 2-fold lower in γδ than in αβ T cells from the patients using different TCR- or CD3-specific mAbs, both in primary (Figure 3B) and in cultured T cells (Supplemental Figure 3B). TCR downregulation after engagement by anti-CD3, however, was similar in both T cell lineages (Supplemental Figure 6A). In contrast, early activation events such as CD69 or CD25 induction were strongly reduced (Figure 3C and Supplemental Figure 6, B and C). αβ and γδ T cells were nevertheless capable of normal anti-CD3– or phytohemagglutinin-induced short-term proliferation on a per-cell basis (5 days; Supplemental Figure 6D). This was confirmed in culture using allogeneic feeder cells (Figure 3D). However, after day 25 in the same cultures, αβ, but not γδ, T cells showed impaired growth relative to a control. Therefore, the CD3D mutation impaired TCR expression and several functions in γδ and αβ T cells, albeit with some contrasting effects: lower TCR expression by γδ T cells but lower in vitro long-term survival of αβ T cells. CD3δ has been reported to bind less strongly to the TCRγδ than to the TCRαβ heterodimer (11), offering a potential mechanism for the observed differential surface levels of TCR in γδ versus αβ T cells when CD3δ is reduced.
The thymus was normal in size at diagnosis (Supplemental Figure 7). To study its function, several studies were performed, including analyses of recent thymic emigrants defined as CD4+CD45RA+CD31+ cells, CD45RA+ (naive) and CD45R0+ (memory) T cells, CD25 expression; TCRB clonality, and TCRVβ usage (Supplemental Figures 8 and 9). The results indicated that the patients’ thymuses produced very few αβ T lymphocytes, and most of these had differentiated into effector memory T cells with an activated phenotype and an oligoclonal TCRVβ repertoire. They may have contributed to the observed in vivo Th2 features in patient AIII.1 (hyper-IgE, eosinophilia, and atopic dermatitis). Similar Th2 immunopathology has been observed in lymphopenic patients with Omenn-like syndrome associated with several primary immunodeficiencies (12) and in mice with partial T cell immunodeficiency (13).
These results indicated that the CD3D splicing mutation strongly impaired αβ, but not γδ, T lymphocyte selection in the thymus, resembling the phenotype of CD3δ-KO mice (14). In contrast, human complete CD3δ immunodeficiencies uniformly lack all T cells (4, 5). Therefore, the leaky mutation, which we believe to be novel, reveals that human αβ and γδ T lymphocytes have differential CD3δ requirements for selection that have not been described for other invariant chains (CD3γ, CD3ε, or TCRζ; Supplemental Table 1). The Tαβ–Tγδ+B+NK+ phenotype has been reported recently in 2 children with TCRα deficiency (3). However, those patients remained relatively healthy for 6–7 years on antibiotic prophylaxis before transplantation; they had activated germinal centers and specific Ab responses against vaccines and autoantigens, likely due to γδ T cell help by homology with the mouse model. The leaky CD3δ SCID patients reported here required very early transplantation (before 1–2 years), lacked germinal centers, and showed very poor T cell–dependent B cell function, perhaps due to the signaling impairment observed also in their γδ T cells (Figure 3C). Thus, CD3δ levels are sufficient for selection, but not for normal function of γδ T cells, as described for CD4+ T cells in Zap70 SCID patients (2).
Two mechanisms may be proposed to explain the selective effect of low CD3δ levels in αβ T lymphocyte development. One is impaired assembly or signaling of the immature precursor of the TCRαβ termed the pre-TCR (15). The second one is impaired signaling through the mature TCRαβ due to reduced interactions of CD3δ with an evolutionarily conserved motif in the TCRα chain membrane-proximal constant region termed the α-chain connecting peptide (16).
Together, the results showed that the leaky CD3D mutation reduced CD3δ chains, which in turn blocked αβ rather than γδ T cell selection. In mature T cells, TCRγδ expression was more impaired than TCRαβ expression, but early signaling through both was similarly impaired.
-
Methods
Further information can be found in Supplemental Methods.
Mutation detection. CD3 amplimers were generated by RT-PCR of PBMC RNA with specific primers (Supplemental Tables 2 and 3). CD3D exons and flanking intronic sequences were amplified from leukocyte DNA by PCR using specific primers and sequenced following standard techniques. Screening for the IVS2+5G→A mutation was performed by RFLP using BsaAI (New England Biolabs). Ecuadorian DNA samples were provided by Antonio Arnaiz-Villena (Complutense University).
Quantitative PCR. TaqMan PCR of PBMC cDNA was done using CD3D WT– or ΔEx2-specific primers and probes (Supplemental Table 4). Samples were normalized to the Ct of CD3E-specific primers and probe 49 from the Universal ProbeLibrary Human (Roche) and to the highest value in each data set, which is shown as 1.
Transfections and Western blotting. 293T cells were transiently transfected using lipofectamine (Invitrogen) with 2 μg of plasmid pIRES-GFP1a (Invitrogen) containing ΔEx2 CD3D cDNA, lysed and analyzed by Western blotting, together with cultured T cells, using APA1/2 (mouse anti-CD3δ cytoplasmic tail IgG mAb; provided by Balbino Alarcón, Centro de Biología Molecular, Madrid, Spain) or M20epsilon (goat anti-CD3ε IgG; Santa Cruz Biotechnology Inc.).
Immunological investigations. Lymphocyte phenotype was determined by flow cytometry using anti-CD3 (Leu4/SK7), anti-CD69 (L78), anti-CD19 (SJ25C1), anti-CD16 (73.1), anti-CD56 (NCAM16.2), and anti-TCRγδ (11F2) from BD Biosciences; and anti-TCRαβ (BMA031 or WT31), anti-TCRγδ (IMMU510), anti-CD4 (13B8.4), and anti-CD8 (B9.11) from Beckman Coulter Immunotech. T cell function was tested by standard overnight H3-thymidine uptake assays (1 μCi/well) by culturing 105 PBMCs with phytohemagglutinin (1 μg/ml; Sigma-Aldrich) or plastic-bound anti-CD3 (10 μg/ml UCHT-1 from BD) for 72 hours. NK cell function was tested by standard 51Cr release assays using PBMCs as effectors and K562 cells as targets. Specific lysis was calculated from cpm as 100 × (sample)/(maximum), after correcting for blank measurements.
T cell lines were generated from PBMCs and expanded weekly with irradiated allogeneic feeder cells (PBMCs and EBV-transformed B cells) at 1:5:5 ratios and final 106 cells/ml in IMDM medium (PAA) with 0.1 μg/ml phytohemagglutinin (only at day 0; Sigma-Aldrich), 40 IU/ml rIL-2 (provided by Craig W. Reynolds, Frederick Cancer Research and Development Center, NCI, NIH, Frederick, Maryland, USA), 10% AB+ human serum, and 1% glutamine (Gibco; Invitrogen).
Study approval. The study was conducted according to the principles expressed in the Declaration of Helsinki and approved by the Hospital Clínico Research Ethics Committee, Madrid, Spain. All participants or their guardians provided informed consent for the collection of samples and subsequent analyses.
Statistics. Bar graph data represent mean ± SD. For proliferation and cytotoxicity, median values of triplicates were used.
-
Acknowledgments
This work was supported by grants from the Spanish NIH (ISCIII — PI080921, PI060057, PI080045, RIER), Ministry of Health (2010 Cohesion Fund for Rare Diseases), Ministry of Science (SAF2011-24235), Fundación Mutua Madrileña, and the Deutsche-Forschungsgemeinschaft (EXC294/BIOSS, SFB620B6). We thank Bruno Hernández, Verónica Pérez-Flores, Alberto C. Guardo, Edgar Fernandez-Malavé, Pilar Perez Breña (National Microbiology Center), and Pieter C.M. Res (University of Amsterdam) for technical support and comments.
Address correspondence to: José R. Regueiro, Inmunología, Facultad de Medicina, Universidad Complutense, 28040 Madrid, Spain. Phone: 34913941642; Fax: 34913941641; E-mail: regueiro@med.ucm.es.
-
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Reference information: J Clin Invest. 2011;121(10):3872–3876. doi:10.1172/JCI44254.
-
References
- Weiss A, Littman DR. Signal transduction by lymphocyte antigen receptors. Cell. 1994;76(2):263–274.
- Notarangelo LD, et al. Primary immunodeficiencies: 2009 update The International Union of Immunological Societies (IUIS) Primary Immunodeficiencies (PID) Expert Committee. J Allergy Clin Immunol. 2009;124(6):1161–1178.
- Morgan NV, et al. Mutation in the TCRα subunit constant gene (TRAC) leads to a human immunodeficiency disorder characterized by a lack of TCRαβ+ T cells. J Clin Invest. 2011;121(2):695–702.
- Dadi HK, Simon AJ, Roifman CM. Effect of CD3delta deficiency on maturation of alpha/beta and gamma/delta T-cell lineages in severe combined immunodeficiency. N Engl J Med. 2003;349(19):1821–1828.
- de Saint Basile G, et al. Severe combined immunodeficiency caused by deficiency in either the delta or the epsilon subunit of CD3. J Clin Invest. 2004;114(10):1512–1517.
- Siegers GM, et al. Different composition of the human and the mouse gammadelta T cell receptor explains different phenotypes of CD3gamma and CD3delta immunodeficiencies. J Exp Med. 2007;204(11):2537–2544.
- Roberts JL, et al. T–B+NK+ severe combined immunodeficiency caused by complete deficiency of the CD3zeta subunit of the T-cell antigen receptor complex. Blood. 2007;109(8):3198–3206.
- Wada T, et al. Detection of T lymphocytes with a second-site mutation in skin lesions of atypical X-linked severe combined immunodeficiency mimicking Omenn syndrome. Blood. 2008;112(5):1872–1875.
- Ehl S, et al. A variant of SCID with specific immune responses and predominance of γδ T cells. J Clin Invest. 2005;115(11):3140–3148.
- Enders A, et al. A severe form of human combined immunodeficiency due to mutations in DNA ligase IV. J Immunol. 2006;176(8):5060–5068.View this article via: PubMed Google Scholar
- Alibaud L, Arnaud J, Llobera R, Rubin B. On the role of CD3d chains in TCRgd/CD3 complexes during assembly and membrane expression. Scand J Immunol. 2001;54(1–2):155–162.
- Ozcan E, Notarangelo LD, Geha RS. Primary immune deficiencies with aberrant IgE production. J Allergy Clin Immunol. 2008;122(6):1054–1062.
- Liston A, Enders A, Siggs OM. Unravelling the association of partial T-cell immunodeficiency and immune dysregulation. Nat Rev Immunol. 2008;8(7):545–558.
- Dave VP, et al. CD3δ deficiency arrests development of the αβ but not the γδ T cell lineage. EMBO J. 1997;16(6):1360–1370.
- Yamasaki S, Saito T. Molecular basis for pre-TCR-mediated autonomous signaling. Trends Immunol. 2007;28(1):39–43.
- Bäckström BT, Müller U, Hausmann B, Palmer E. Positive selection through a motif in the alphabeta T cell receptor. Science. 1998;281(5378):835–838.
-
Version history
- Version 1 (September 19, 2011): No description
- Version 2 (October 3, 2011): No description



Copyright © 2026 American Society for Clinical Investigation
ISSN: 0021-9738 (print), 1558-8238 (online)